Lida Wang
@LD_matrix
Postdoc @DanaFarber and @harvardmed | Biostat Ph.D @penn_state | Statistical genetics, functional genomics, data science
Bạn có thể thích
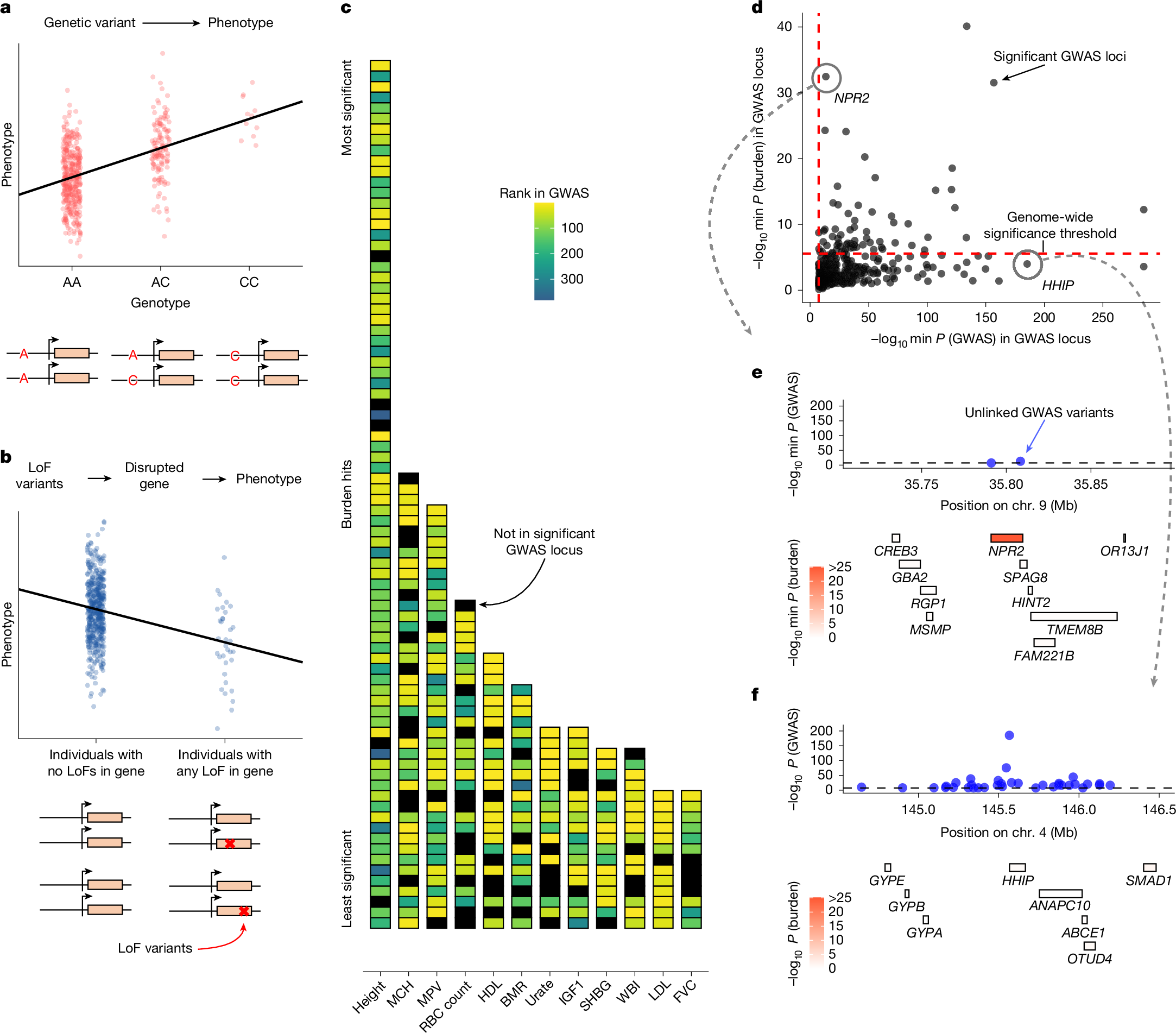
How do GWAS and rare variant burden tests rank gene signals? In new work @Nature with Jeff Spence, @jkpritch, and our wonderful coauthors we find the key factors are what we call Specificity, Length, and Luck! 🧬🧪🧵 nature.com/articles/s4158…
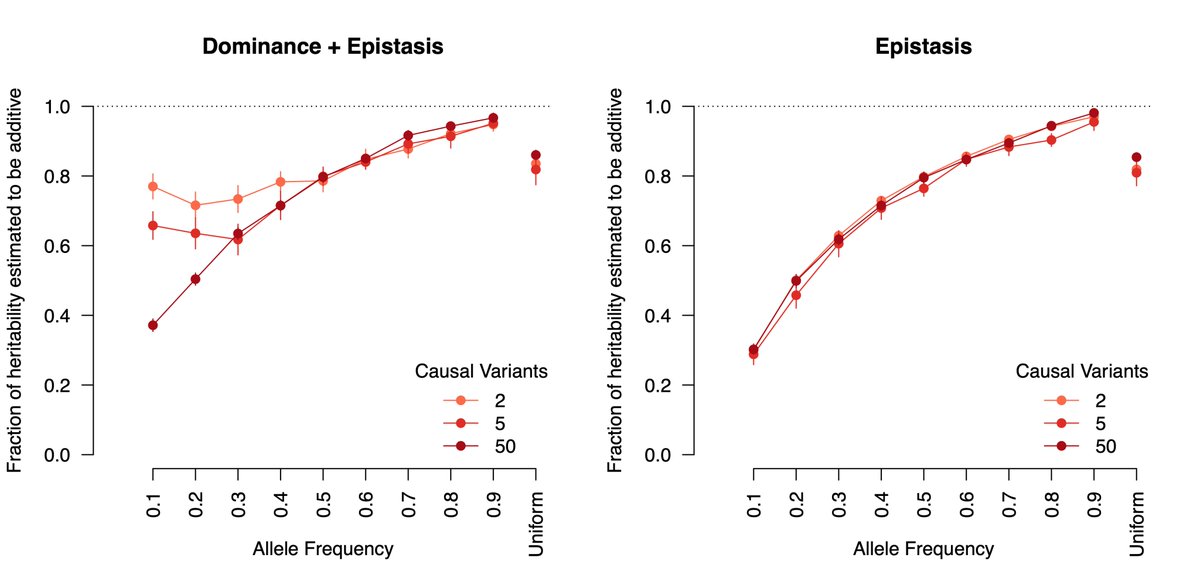
The proportion of epistatic heritability that is estimated as additive by quantitative genetic models. Epistasis deviates more from additivity for lower frequency causal alleles, but on average >80% of biological GxG will just look like statistical G.
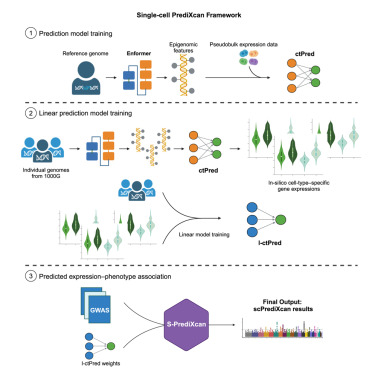
Check out our scPrediXcan paper cell.com/cell-genomics/… Led by talented @Charles_Zhou12, supervised by @MengjieChen6 and me, with thanks to many contributors scPrediXcan integrates deep learning and single cell expression data into a powerful cell type specific TWAS framework.
I'm excited to share a wonderful collaboration with @LD_matrix: JOBS integrates bulk and single-cell eQTLs to boost eQTL discovery. Integrated with GWAS, it identified more disease-linked loci for 14 immune disorders & fueled drug-repurposing. #immunology #drugdiscovery #genomics
I am excited to share our new work that generates a single cell eQTL atlas for immune cell types. The work integrates bulk eQTLs with sc-eQTLs, which boosts the power for identifying sc-eQTLs for up to 4 folds. As such, we have the power equivalent to sc-RNASeq of 4K individuals.
🧠🔥"Put on your 3D glasses...oh wait, you don't need them!"😝 Brain Metabolome just got an upgrade-why look at slices when you can see the whole picture in 3D? Check out our latest work in @NatMetabolism .🚀🔬 🔗doi.org/10.1038/s42255…

Beyond excited to share this new paper with all of you . It's the most fun we've ever had. We figured out how to study a latent index driving partner choice without measuring it directly🥂 @qinwen_zzz Preprint📰: biorxiv.org/content/10.110… Sumstats⬇️: qlu-lab.org/data.html

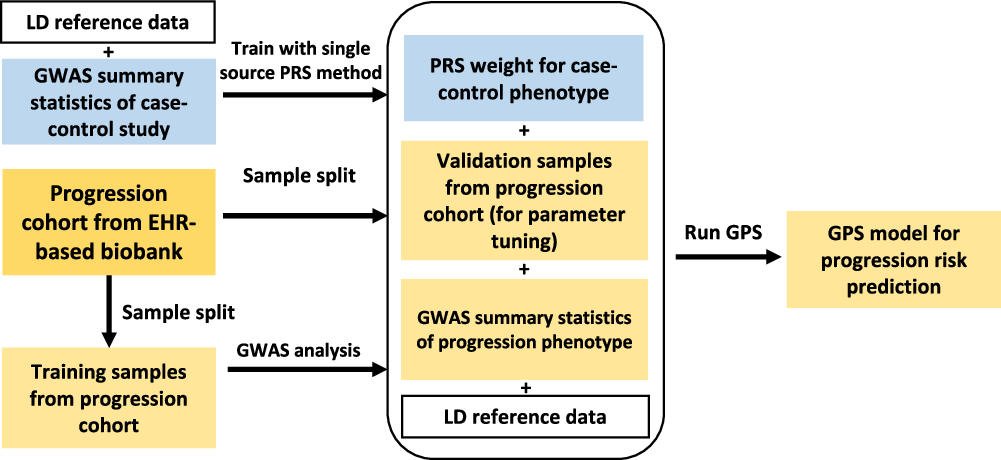
I am happy to share our new work that predicts the progression of autoimmune diseases, integrating EHR and GWAS datasets @NatureComms . nature.com/articles/s4146…
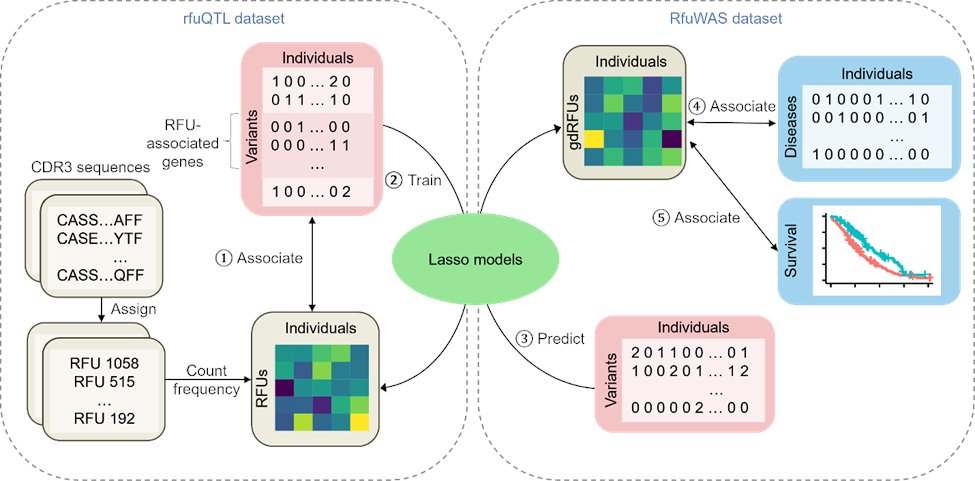
Excited to share our recent publication in Communications Biology! We explored how genetic variation shapes T cell receptor (TCR) diversity and its impact on diseases such as autoimmune disorders and cancer. nature.com/articles/s4200… (1/11)
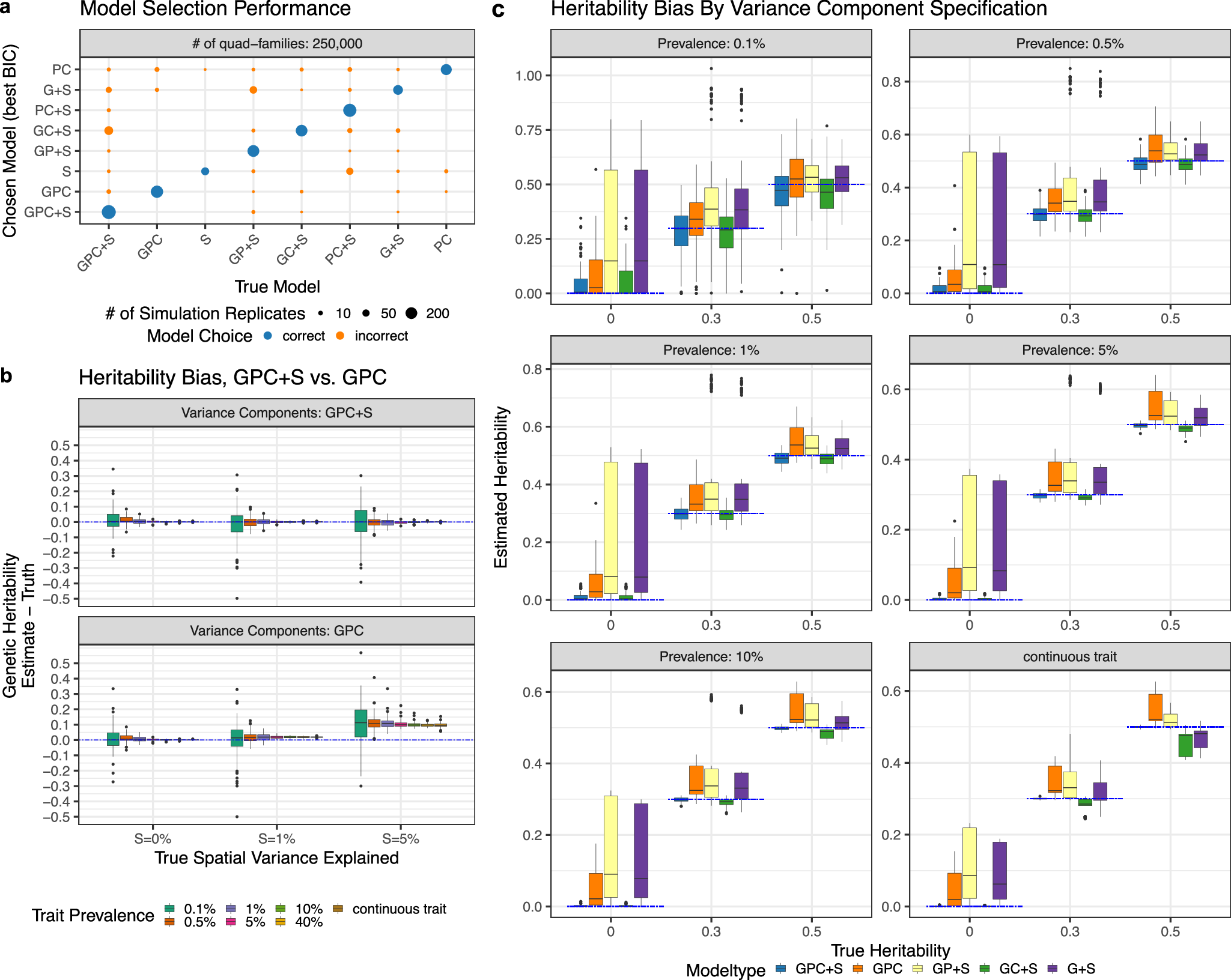
Thrilled to share our work to analyze insurance claim datasets to dissect genetic and environmental contributions to 1083 human diseases. We developed a spatial linear mixed model SMILE, using geolocations as a proxy for the environment. nature.com/articles/s4146…
nature.com
Dissecting heritability, environmental risk, and air pollution causal effects using > 50 million...
Nature Communications - Large national-level electronic health record datasets offer new opportunities for disentangling the roles of genes and environment in human diseases. Here, the authors...
Excited to share our latest work describing a spatial mixed linear effect (SMILE) model that refines the estimates of genetic heritability and air pollution causal effects using EHR in @NatureComms. Huge thanks to Dan McGuire and my PI @dajiangliu81! nature.com/articles/s4146…
Really excited to share our latest work in @NatureComms! We developed a method called EXPRESSO to conduct TWAS using single-cell eQTL summary statistics. Special thanks to @dajiangliu81 for the guidance and the help from @GUTomics and lab members.
Delighted to share our work that integrates single cell eQTL data with GWAS to identify cell type specific risk genes for complex traits. Published today at @NatureComms . Work led by talented students @LD_matrix and @GUTomics . nature.com/articles/s4146…
Excited to share our latest work in @NatureComms. We extended our previous PUMICE's framework (nature.com/articles/s4146…) to allow the utilization of publicly available single-cell eQTL summary statistics. Kudos to @LD_matrix and @dajiangliu81 for leading this important work.
Delighted to share our work that integrates single cell eQTL data with GWAS to identify cell type specific risk genes for complex traits. Published today at @NatureComms . Work led by talented students @LD_matrix and @GUTomics . nature.com/articles/s4146…
Hi friends! We are looking to hire a postdoc to join our group @CMUCompBio @SCSatCMU, to work on exciting projects related to identifying disease genes by integrating GWAS and functional genomics. mzhanglab.github.io/openings/
Excited to share our latest work (nature.com/articles/s4146…) in @NatureComms. We conducted TWAS focusing on blood pressure traits using PUMICE, complemented by multi-omics follow-up analyses. Kudos to Xiaoguang Xu, @EalesJames, and Maciej Tomaszewski for leading this amazing work.
It's 2024 and there are many great tutorials for single-cell analysis. One frustrating thing to follow any R-based tutorial is package installation (of course)! With all sorts pkg managers, it should be easy to install pkgs, right? Well, no, especially if you are a beginner. 1/9
A few thoughts on the recent set of papers torture testing genomic deep learning for predicting individual-level gene expression [ pubmed.ncbi.nlm.nih.gov/38036790/ , pubmed.ncbi.nlm.nih.gov/38036778/ , pubmed.ncbi.nlm.nih.gov/38036789/ ]. First a brief summary 🧵:
Links to the recordings: Day 1: nih.zoomgov.com/rec/play/9o8-x… nih.zoomgov.com/rec/play/YZutj… Day 2: nih.zoomgov.com/rec/play/V1meR… nih.zoomgov.com/rec/play/tr_e8…
Enjoying re-watching Day 1 of this NHGRI meeting on genetic architecture [genome.gov/event-calendar…]. My notes below, with comments in [].
United States Xu hướng
- 1. Florida 100K posts
- 2. Texas 171K posts
- 3. #SmallBusinessSaturday 2,238 posts
- 4. Ohio State 26.8K posts
- 5. Kentucky 14.2K posts
- 6. Buckeyes 4,641 posts
- 7. Go Blue 6,487 posts
- 8. Tyler Adams 2,542 posts
- 9. Good Saturday 35.1K posts
- 10. Go Bucks 2,158 posts
- 11. Raphinha 17.4K posts
- 12. #MeAndTheeSeriesEP3 1.39M posts
- 13. Saban 5,142 posts
- 14. Gameday 29.2K posts
- 15. Bernal 13.9K posts
- 16. #SaturdayVibes 3,860 posts
- 17. #StrayKidsAtMAMA2025 12.7K posts
- 18. #Caturday 3,666 posts
- 19. The Game 960K posts
- 20. Nunes 23.3K posts
Something went wrong.
Something went wrong.