#noncodingvariants search results
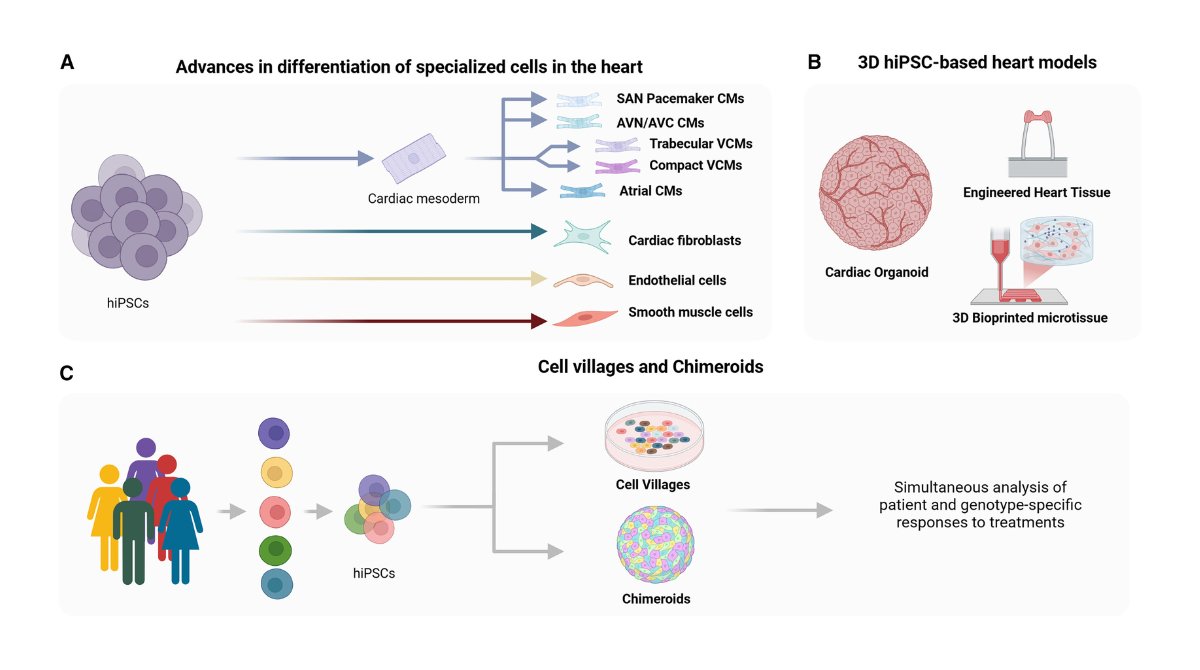
Online now! Review discussing the basics of noncoding variant-mediated pathogenesis, the methodologies utilized, and how #iPSC -based heart models can be leveraged to dissect the mechanisms of #noncodingvariants ow.ly/wqcH50VlLGC @UBCcps @BCCHresearch @SBME_UBC @ISSCR
Annotating and prioritizing human non-coding variants with RegulomeDB v.2. #NonCodingVariants #VariantsAnnotation #VariantsPrioritization @NatureGenet nature.com/articles/s4158…
nature.com
Annotating and prioritizing human non-coding variants with RegulomeDB v.2
Nature Genetics - Annotating and prioritizing human non-coding variants with RegulomeDB v.2
Recommendations for clinical interpretation of variants found in non-coding regions of the genome. #NonCodingVariants #Clinicalinterpretation @j_ellingford @nickywhiffin @GenomeMedicine genomemedicine.biomedcentral.com/articles/10.11…
New Topic "#NonCodingVariants: From #Interpretation to #PersonalizedMedicine" led by Drs Elvezia Maria Paraboschi @HumanitasMilano, Elisabetta Morini and Harrison Brand @Harvard and @hongheeW is open for submissions. Submit here: fro.ntiers.in/3oa7

Classification of non-coding variants with high pathogenic impact. #WGS #NonCodingVariants #FunctionalPrediction #DiseaseImpact #MachineLearning #Bioinformatics journals.plos.org/plosgenetics/a… @PLOSGenetics
Cell type–specific interpretation of noncoding variants using deep learning–based methods. #NonCodingVariants #DeepLearning #CellTypeVariants @GigaScience academic.oup.com/gigascience/ar…
academic.oup.com
Cell type–specific interpretation of noncoding variants using deep learning–based methods
Abstract. Interpretation of noncoding genomic variants is one of the most important challenges in human genetics. Machine learning methods have emerged rec
Non-coding variants impact cis-regulatory coordination in a cell type-specific manner. #NonCodingVariants #CisRegulatoryElements @GenomeBiology genomebiology.biomedcentral.com/articles/10.11…
“By using aberrant #genetranscription to reveal the function of #noncodingvariants, we developed cis-X to enable discovery of noncoding variants driving #cancer in individual tumor #genomes" - @StJudeResearch rna-seqblog.com/st-jude-resear…
Performance Comparison of Computational Prediction Methods for the Function and Pathogenicity of Non-coding Variants. #NonCodingVariants #FunctionPrediction #ToolsBenchmarking biorxiv.org/content/10.110… @biorxivpreprint #Bioinformatics
A framework for detecting noncoding rare variant associations of large-scale whole-genome sequencing studies. #NonCodingVariants #WGSdata #Bioinformatics biorxiv.org/content/10.110…
Structural and non-coding variants increase the diagnostic yield of clinical whole genome sequencing for rare diseases. #StructuralVariants #SVs #NoncodingVariants #Diagnostic #WGS #RareDiseases @GenomeMedicine genomemedicine.biomedcentral.com/articles/10.11…
Classification of non-coding variants with high pathogenic impact. #NonCodingVariants #PathogenicityPrioritization journals.plos.org/plosgenetics/a… @PLOSGenetics
TVAR: Assessing Tissue-specific Functional Effects of Non-coding Variants with Deep Learning. #NonCodingVariants #DeepLearning #Bioinformatics academic.oup.com/bioinformatics…
academic.oup.com
TVAR: assessing tissue-specific functional effects of non-coding variants with deep learning
AbstractMotivation. Analysis of whole-genome sequencing (WGS) for genetics is still a challenge due to the lack of accurate functional annotation of non-co
WEVar: a novel statistical learning framework for predicting noncoding regulatory variants. #NonCodingVariants academic.oup.com/bib/advance-ar… @BriefingBioinfo
Great talk from @j_ellingford on interpreting #NonCodingVariants in @GenomicsEngland data with focus in ophthalmological disorders #LGN2022 @LdnGeneNet
Reclutada durante años en Dpto. Genética @Hospital_FJD, el Servicio de Oftalmología del HU Clínico San Carlos y Fac Med Albacete. Esperamos además mejorar las tasas de caracterización genética 🧬, identificar nuevos genes, implicación #noncodingvariants 🧵
The Impact of Rare Noncoding Variants scienceness.com/news/6667-the-… #Scienceness #TranscriptomeSequencing #NoncodingVariants #eQTLs #sQTLs
Great talk from @j_ellingford on interpreting #NonCodingVariants in #RareDisease 🧬 Highlighted available New guidelines for standardized clinical variant interpretation & that Functional information & methods used are critical for variant interpretation. #LGN2022 @LdnGeneNet
#BankaGrpJC 03_2020 Journal Club on #Zoom! Last week we discussed this interesting article on #NoncodingVariants #SNPs rdcu.be/b3qej
“Predicting the impact of #noncodingvariants is still a challenge” @alexisjbattle #WGS & #RNA-seq data GTEx #bog16 genomeweb.com/sequencing-tec…
Online now! Review discussing the basics of noncoding variant-mediated pathogenesis, the methodologies utilized, and how #iPSC -based heart models can be leveraged to dissect the mechanisms of #noncodingvariants ow.ly/wqcH50VlLGC @UBCcps @BCCHresearch @SBME_UBC @ISSCR

Reclutada durante años en Dpto. Genética @Hospital_FJD, el Servicio de Oftalmología del HU Clínico San Carlos y Fac Med Albacete. Esperamos además mejorar las tasas de caracterización genética 🧬, identificar nuevos genes, implicación #noncodingvariants 🧵
Non-coding variants impact cis-regulatory coordination in a cell type-specific manner. #NonCodingVariants #CisRegulatoryElements @GenomeBiology genomebiology.biomedcentral.com/articles/10.11…
Structural and non-coding variants increase the diagnostic yield of clinical whole genome sequencing for rare diseases. #StructuralVariants #SVs #NoncodingVariants #Diagnostic #WGS #RareDiseases @GenomeMedicine genomemedicine.biomedcentral.com/articles/10.11…
Annotating and prioritizing human non-coding variants with RegulomeDB v.2. #NonCodingVariants #VariantsAnnotation #VariantsPrioritization @NatureGenet nature.com/articles/s4158…
nature.com
Annotating and prioritizing human non-coding variants with RegulomeDB v.2
Nature Genetics - Annotating and prioritizing human non-coding variants with RegulomeDB v.2
Cell type–specific interpretation of noncoding variants using deep learning–based methods. #NonCodingVariants #DeepLearning #CellTypeVariants @GigaScience academic.oup.com/gigascience/ar…
academic.oup.com
Cell type–specific interpretation of noncoding variants using deep learning–based methods
Abstract. Interpretation of noncoding genomic variants is one of the most important challenges in human genetics. Machine learning methods have emerged rec
Great talk from @j_ellingford on interpreting #NonCodingVariants in @GenomicsEngland data with focus in ophthalmological disorders #LGN2022 @LdnGeneNet
Great talk from @j_ellingford on interpreting #NonCodingVariants in #RareDisease 🧬 Highlighted available New guidelines for standardized clinical variant interpretation & that Functional information & methods used are critical for variant interpretation. #LGN2022 @LdnGeneNet
TVAR: Assessing Tissue-specific Functional Effects of Non-coding Variants with Deep Learning. #NonCodingVariants #DeepLearning #Bioinformatics academic.oup.com/bioinformatics…
academic.oup.com
TVAR: assessing tissue-specific functional effects of non-coding variants with deep learning
AbstractMotivation. Analysis of whole-genome sequencing (WGS) for genetics is still a challenge due to the lack of accurate functional annotation of non-co
Recommendations for clinical interpretation of variants found in non-coding regions of the genome. #NonCodingVariants #Clinicalinterpretation @j_ellingford @nickywhiffin @GenomeMedicine genomemedicine.biomedcentral.com/articles/10.11…
Classification of non-coding variants with high pathogenic impact. #WGS #NonCodingVariants #FunctionalPrediction #DiseaseImpact #MachineLearning #Bioinformatics journals.plos.org/plosgenetics/a… @PLOSGenetics
Classification of non-coding variants with high pathogenic impact. #NonCodingVariants #PathogenicityPrioritization journals.plos.org/plosgenetics/a… @PLOSGenetics
A framework for detecting noncoding rare variant associations of large-scale whole-genome sequencing studies. #NonCodingVariants #WGSdata #Bioinformatics biorxiv.org/content/10.110…
Performance Comparison of Computational Prediction Methods for the Function and Pathogenicity of Non-coding Variants. #NonCodingVariants #FunctionPrediction #ToolsBenchmarking biorxiv.org/content/10.110… @biorxivpreprint #Bioinformatics
WEVar: a novel statistical learning framework for predicting noncoding regulatory variants. #NonCodingVariants academic.oup.com/bib/advance-ar… @BriefingBioinfo
Classification of non-coding variants with high pathogenic impact. #WGS #NonCodingVariants #FunctionalPrediction #DiseaseImpact #MachineLearning #Bioinformatics biorxiv.org/content/10.110…
New Topic "#NonCodingVariants: From #Interpretation to #PersonalizedMedicine" led by Drs Elvezia Maria Paraboschi @HumanitasMilano, Elisabetta Morini and Harrison Brand @Harvard and @hongheeW is open for submissions. Submit here: fro.ntiers.in/3oa7

“By using aberrant #genetranscription to reveal the function of #noncodingvariants, we developed cis-X to enable discovery of noncoding variants driving #cancer in individual tumor #genomes" - @StJudeResearch rna-seqblog.com/st-jude-resear…
#BankaGrpJC 03_2020 Journal Club on #Zoom! Last week we discussed this interesting article on #NoncodingVariants #SNPs rdcu.be/b3qej
“Predicting the impact of #noncodingvariants is still a challenge” @alexisjbattle #WGS & #RNA-seq data GTEx #bog16 genomeweb.com/sequencing-tec…
Online now! Review discussing the basics of noncoding variant-mediated pathogenesis, the methodologies utilized, and how #iPSC -based heart models can be leveraged to dissect the mechanisms of #noncodingvariants ow.ly/wqcH50VlLGC @UBCcps @BCCHresearch @SBME_UBC @ISSCR

New Topic "#NonCodingVariants: From #Interpretation to #PersonalizedMedicine" led by Drs Elvezia Maria Paraboschi @HumanitasMilano, Elisabetta Morini and Harrison Brand @Harvard and @hongheeW is open for submissions. Submit here: fro.ntiers.in/3oa7

Something went wrong.
Something went wrong.
United States Trends
- 1. Michigan 143K posts
- 2. Mateer 4,388 posts
- 3. Ohio State 64.7K posts
- 4. Buckeyes 20.8K posts
- 5. Arbuckle N/A
- 6. Tim Banks N/A
- 7. Underwood 10.4K posts
- 8. Ryan Day 9,922 posts
- 9. Vandy 7,907 posts
- 10. Miami 64.8K posts
- 11. Oklahoma 23.6K posts
- 12. Baugh 1,450 posts
- 13. Venezuela 464K posts
- 14. #GoBucks 13.1K posts
- 15. Hawkins 12.7K posts
- 16. #SurvivorSeries 20.7K posts
- 17. Sherrone 5,816 posts
- 18. Pavia 2,762 posts
- 19. #Sooners 1,507 posts
- 20. Stoops 6,493 posts










