#stringdb результаты поиска
@larsjuhljensen is there any way where i can get to know the evidence of interaction when i import data from #STRINGdb using #StringApp in @cytoscape ?
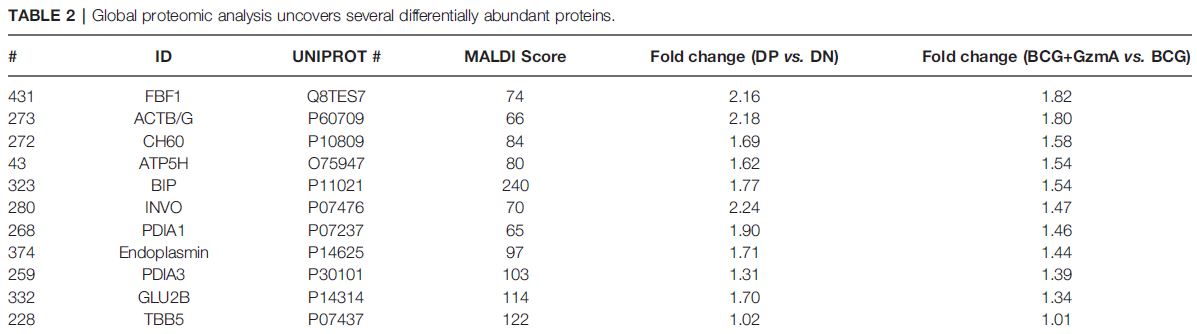
6/10 To better understand the role of GzmA, we performed a global proteomic analysis using 2D-DIGE, where we showed that there are 10 differentially abundant proteins. Using #STRINGDB, we identified ER stress response and ATP production as two important pathways.
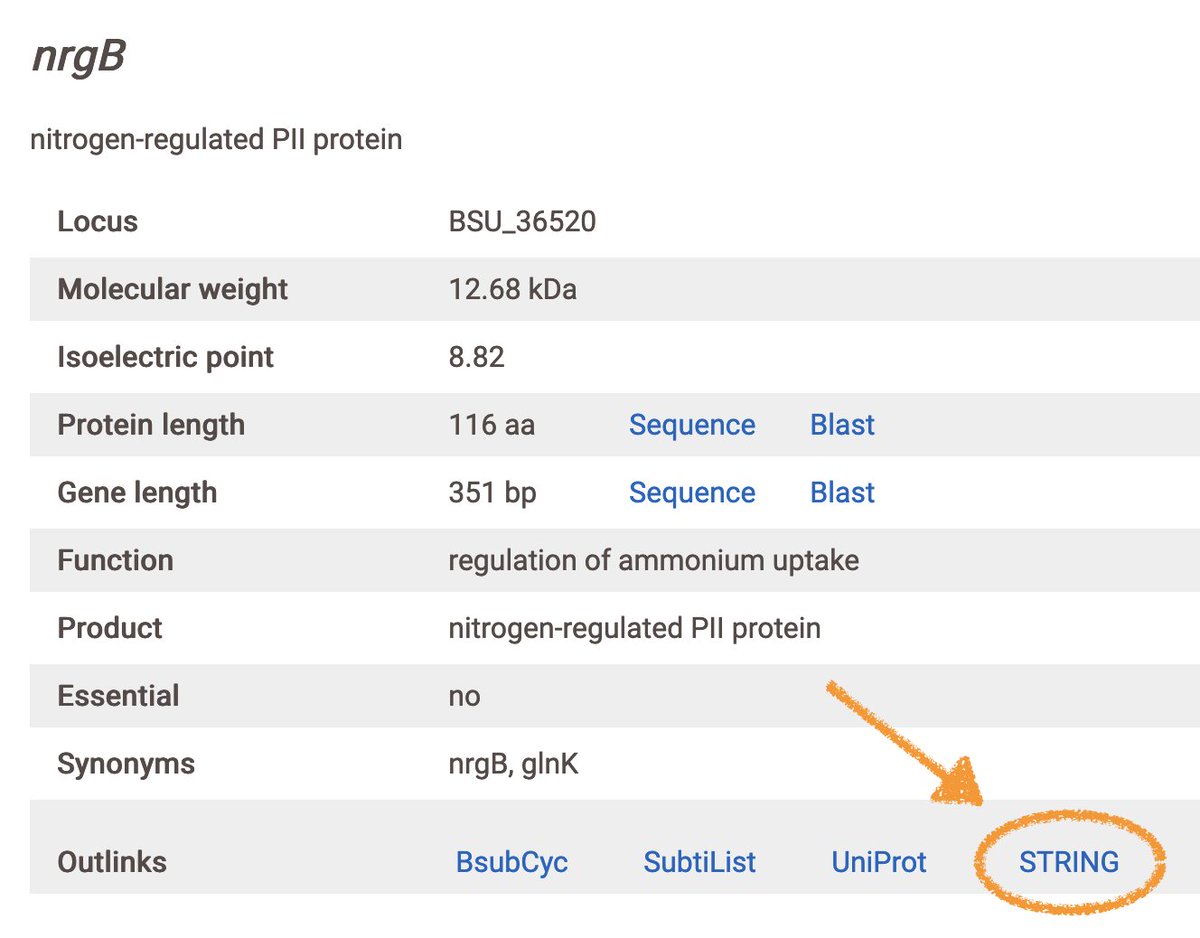
We've added outlinks for string-db.org to subtiwiki.uni-goettingen.de and mycowiki.uni-goettingen.de! Use it to view known and predicted PPIs in an interactive network graph. Example shown for nrgB/glnK: subtiwiki.uni-goettingen.de/v4/gene?id=EC7… #STRINGdb #SubtiWiki #MycoWiki

#STRINGdb Ques: Are you aware of any package which provides different sources of evidence of interaction and associated score from the database? Other than playing with its web interface. At least, I do not see it possible through @Bioconductor #STRINGdb #rstats📦#Bioinformatics

Gathering new info using #stringdb over at http://string-db.org/ and the android tool that linked to it #BiochemistryLabSuite #diybio
This is great! Looking forward to someone combining this tool with proteomics and PPI databases for abundance and interactions, with scRNA-seq for cell-type specificity, and @ProteinAtlas for subcellular localization, to make a virtual cell/multicellular organism.#STRINGdb #VR
#stringDB NW red. strategy: Filter "high confidence (>0.7) interactions", then on 'escore' being part of the combined score @larsjuhljensen?
Glad to share our preprint: SPACE: STRING Proteins As Complementary Embeddings: biorxiv.org/content/10.110… Great thanks to all the contributors: Damian Szklarczyk, Christian von Mering, and @larsjuhljensen #bioinformatics #proteinEmbeddings #STRINGdb #computationalBiology
SPACE: STRING Proteins as Complementary Embeddings • Presenting SPACE: a collection of embeddings that aligns protein network and sequence data across 1,322 eukaryotic species in the STRING database. • Key innovation: SPACE uses the FedCoder framework to align

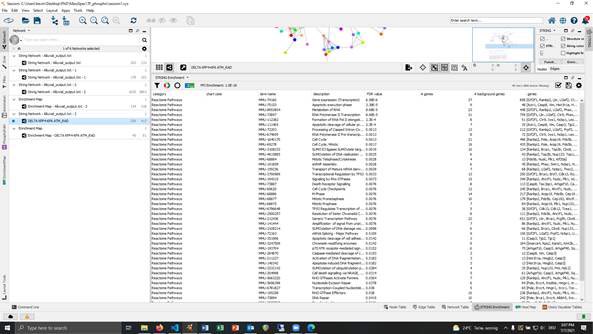
@cytoscape #stringdb since some update in the past month I am not able to reproduce any pathway enrichments made with the String-Plugin in cytoscape. Here are two pictures from the very same dataset I saved beginning of the year. The shorter list is the new calculation.

I tried multiple tools (frequently updated, provides easier access and gives scope to play with required options): g:profiler (@ELIXIREstonia), clusterProfiler (@guangchuangyu), GREAT, @cytoscape, #STRINGdb However, we mostly need post-processing for effective interpretation.
Tweet on #BET1L: not in the same gene network as #STK26 according to #StringDB: string-db.org/network/9606.E… Not linked: string-db.org/cgi/network?ta…
E.g. #BET1L: "Bet1 golgi vesicular membrane trafficking protein like". Very little known about it. Not even a 3D structure in PDBe. No known paralogues, but highly conserved down to the #Chordates 676Mya ensembl.org/Homo_sapiens/G…
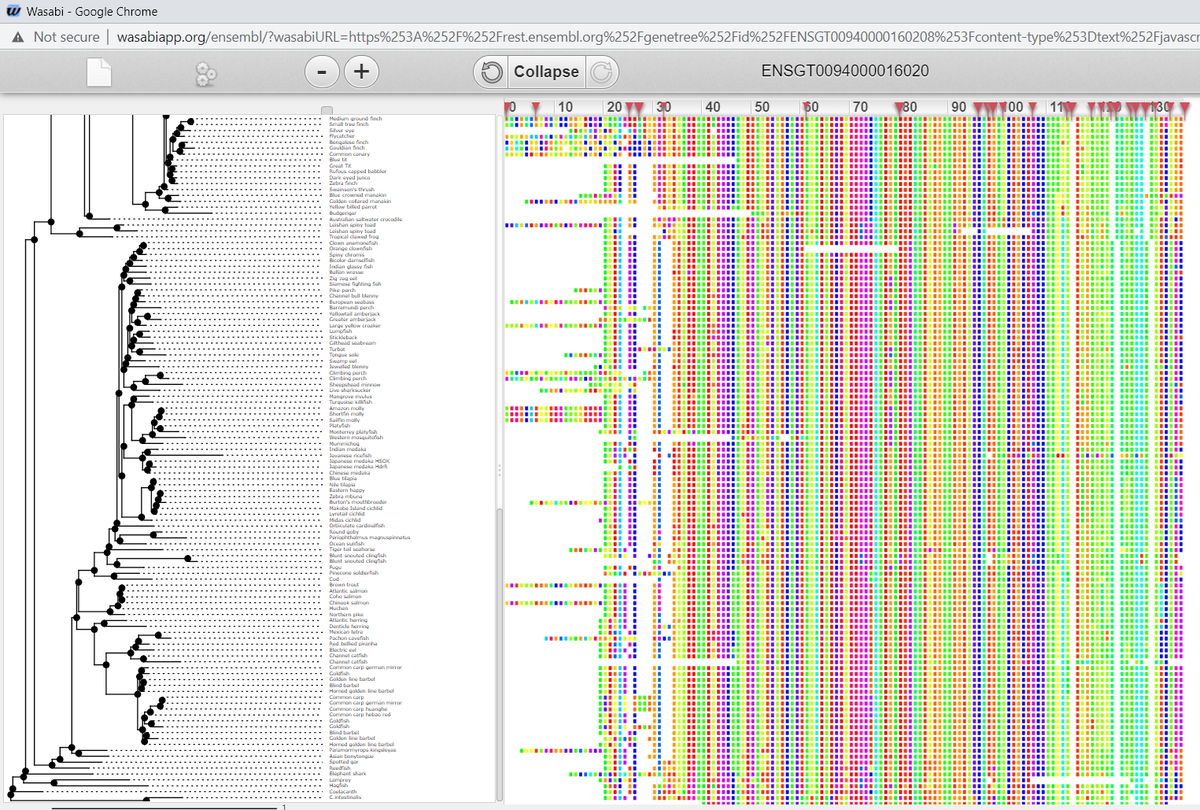
Today we share 64 embedding of #STRINGdb's Species Tree (~20k nodes) from 🍇, including methods integrated from @benrozemberczki's KarateClub and @keenuniverse. The graph is a tree, and that makes some methods go a bit crazy. See the code here: github.com/AnacletoLAB/gr…

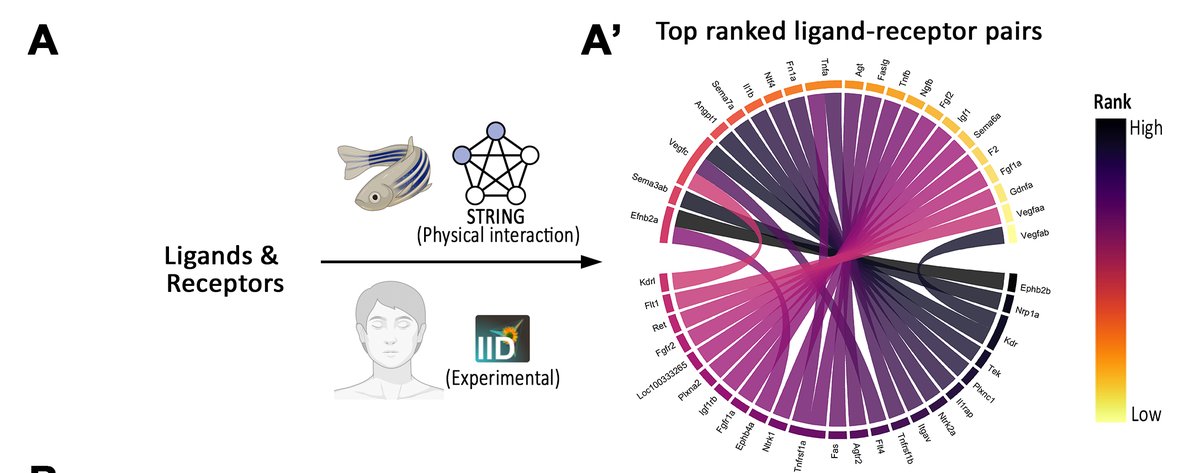
We used #STRINGdb to build a zebrafish-specific ligand-receptor interactome database. We also added human interactions from IID database from Jurisica group 12/n
Glad to share our preprint: SPACE: STRING Proteins As Complementary Embeddings: biorxiv.org/content/10.110… Great thanks to all the contributors: Damian Szklarczyk, Christian von Mering, and @larsjuhljensen #bioinformatics #proteinEmbeddings #STRINGdb #computationalBiology
SPACE: STRING Proteins as Complementary Embeddings • Presenting SPACE: a collection of embeddings that aligns protein network and sequence data across 1,322 eukaryotic species in the STRING database. • Key innovation: SPACE uses the FedCoder framework to align

We used #STRINGdb to build a zebrafish-specific ligand-receptor interactome database. We also added human interactions from IID database from Jurisica group 12/n

We've added outlinks for string-db.org to subtiwiki.uni-goettingen.de and mycowiki.uni-goettingen.de! Use it to view known and predicted PPIs in an interactive network graph. Example shown for nrgB/glnK: subtiwiki.uni-goettingen.de/v4/gene?id=EC7… #STRINGdb #SubtiWiki #MycoWiki


Today we share 64 embedding of #STRINGdb's Species Tree (~20k nodes) from 🍇, including methods integrated from @benrozemberczki's KarateClub and @keenuniverse. The graph is a tree, and that makes some methods go a bit crazy. See the code here: github.com/AnacletoLAB/gr…

I tried multiple tools (frequently updated, provides easier access and gives scope to play with required options): g:profiler (@ELIXIREstonia), clusterProfiler (@guangchuangyu), GREAT, @cytoscape, #STRINGdb However, we mostly need post-processing for effective interpretation.
6/10 To better understand the role of GzmA, we performed a global proteomic analysis using 2D-DIGE, where we showed that there are 10 differentially abundant proteins. Using #STRINGDB, we identified ER stress response and ATP production as two important pathways.

This is great! Looking forward to someone combining this tool with proteomics and PPI databases for abundance and interactions, with scRNA-seq for cell-type specificity, and @ProteinAtlas for subcellular localization, to make a virtual cell/multicellular organism.#STRINGdb #VR
@cytoscape #stringdb since some update in the past month I am not able to reproduce any pathway enrichments made with the String-Plugin in cytoscape. Here are two pictures from the very same dataset I saved beginning of the year. The shorter list is the new calculation.


Tweet on #BET1L: not in the same gene network as #STK26 according to #StringDB: string-db.org/network/9606.E… Not linked: string-db.org/cgi/network?ta…
E.g. #BET1L: "Bet1 golgi vesicular membrane trafficking protein like". Very little known about it. Not even a 3D structure in PDBe. No known paralogues, but highly conserved down to the #Chordates 676Mya ensembl.org/Homo_sapiens/G…

#STRINGdb Ques: Are you aware of any package which provides different sources of evidence of interaction and associated score from the database? Other than playing with its web interface. At least, I do not see it possible through @Bioconductor #STRINGdb #rstats📦#Bioinformatics

@larsjuhljensen is there any way where i can get to know the evidence of interaction when i import data from #STRINGdb using #StringApp in @cytoscape ?
#stringDB NW red. strategy: Filter "high confidence (>0.7) interactions", then on 'escore' being part of the combined score @larsjuhljensen?
Gathering new info using #stringdb over at http://string-db.org/ and the android tool that linked to it #BiochemistryLabSuite #diybio
#STRINGdb Ques: Are you aware of any package which provides different sources of evidence of interaction and associated score from the database? Other than playing with its web interface. At least, I do not see it possible through @Bioconductor #STRINGdb #rstats📦#Bioinformatics

Today we share 64 embedding of #STRINGdb's Species Tree (~20k nodes) from 🍇, including methods integrated from @benrozemberczki's KarateClub and @keenuniverse. The graph is a tree, and that makes some methods go a bit crazy. See the code here: github.com/AnacletoLAB/gr…

We've added outlinks for string-db.org to subtiwiki.uni-goettingen.de and mycowiki.uni-goettingen.de! Use it to view known and predicted PPIs in an interactive network graph. Example shown for nrgB/glnK: subtiwiki.uni-goettingen.de/v4/gene?id=EC7… #STRINGdb #SubtiWiki #MycoWiki


We used #STRINGdb to build a zebrafish-specific ligand-receptor interactome database. We also added human interactions from IID database from Jurisica group 12/n

6/10 To better understand the role of GzmA, we performed a global proteomic analysis using 2D-DIGE, where we showed that there are 10 differentially abundant proteins. Using #STRINGDB, we identified ER stress response and ATP production as two important pathways.

@cytoscape #stringdb since some update in the past month I am not able to reproduce any pathway enrichments made with the String-Plugin in cytoscape. Here are two pictures from the very same dataset I saved beginning of the year. The shorter list is the new calculation.


Something went wrong.
Something went wrong.
United States Trends
- 1. Red Cross N/A
- 2. Pope N/A
- 3. DoorDash N/A
- 4. Catholic N/A
- 5. Leeds N/A
- 6. Ugarte N/A
- 7. Yoro N/A
- 8. Carrick N/A
- 9. #MUFC N/A
- 10. #MUNLEE N/A
- 11. #OurFutureIsPerfectWITH7 N/A
- 12. Martinez N/A
- 13. Manchester United N/A
- 14. Vatican N/A
- 15. Dr. Trump N/A
- 16. #BELIFT_Boycott_Continues N/A
- 17. No Tax N/A
- 18. Casemiro N/A
- 19. Ruby Rose N/A
- 20. McLovin N/A