#slayerstructure search results
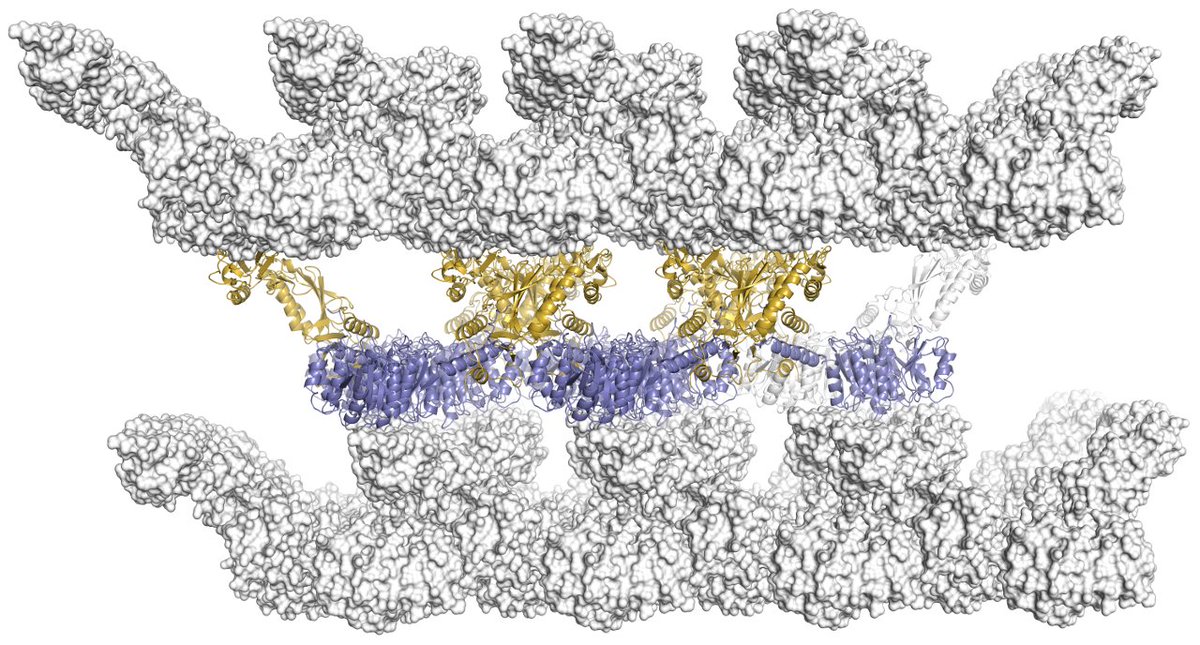
3. The 3D crystals are made of stacks of 2D lattices of SlpA #xraycrystallography #SlayerStructure #BugSlayer 5/n
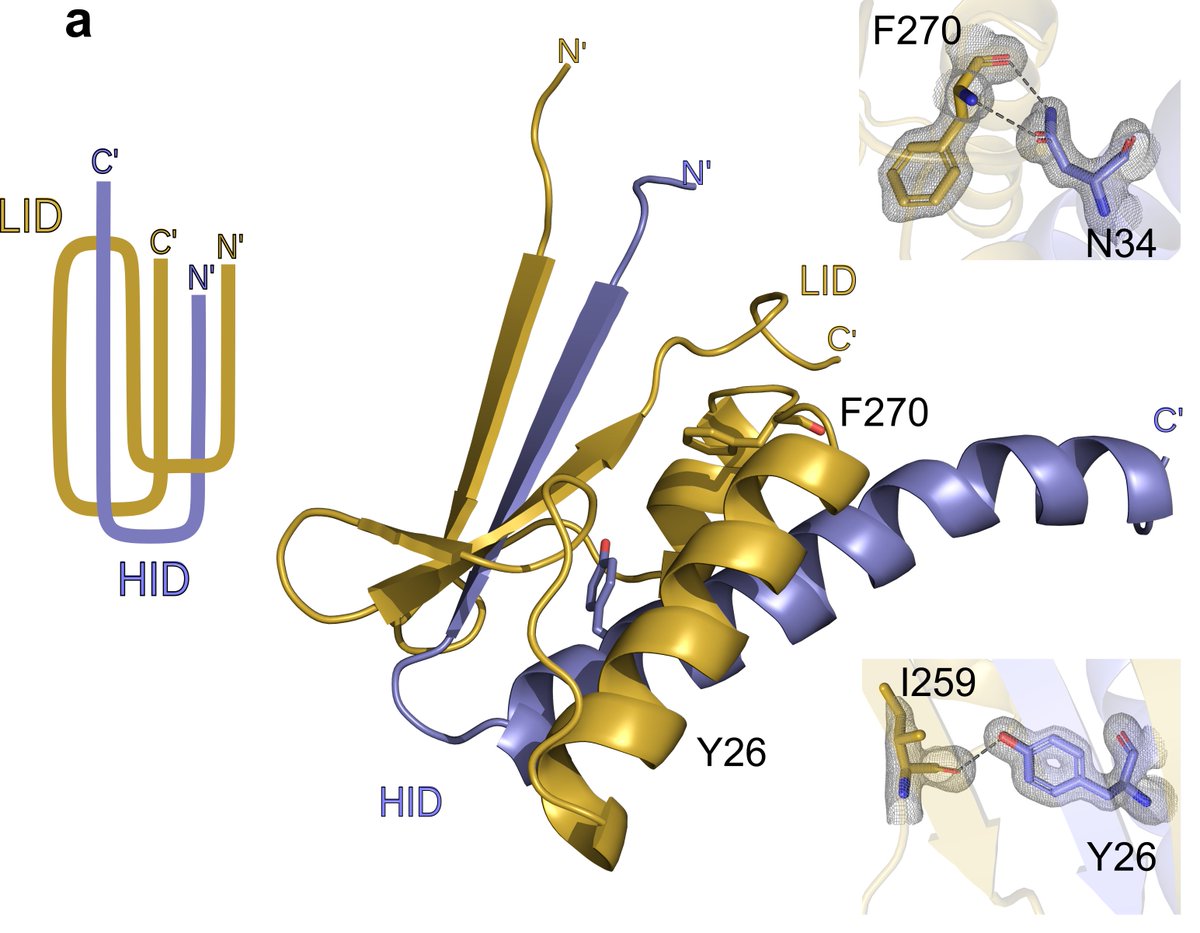
2. the complex is maintained by an intricate “paperclip” motif by the interacting domains. #SlayerStructure #xraycrystallography #BugSlayer 4/n
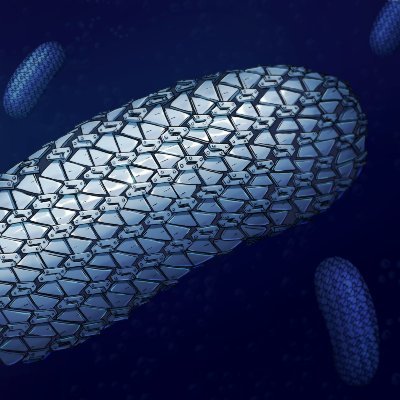
7. The #Cdiff #BugSlayer is very tightly packed, with pores of only about 11A, so only small ions can diffuse through #SlayerStructure 9/n
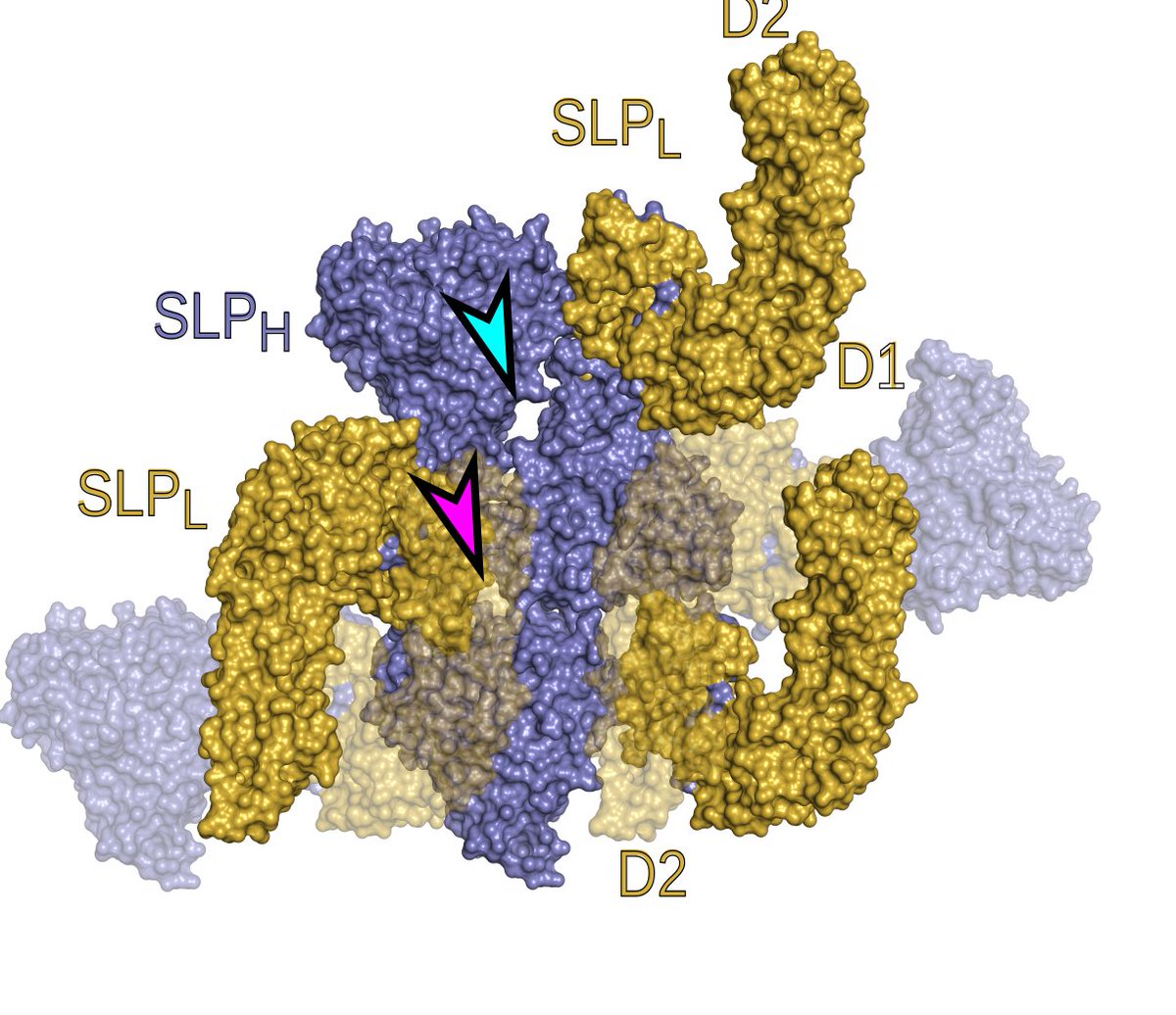
5. The crystal packing of SlpA mimics the assembly of the S-layer in the cells, so interactions in the crystal are a model for how the #BugSlayer assembles. #SlayerStructure 7/n
8. Removing the most exposed SLPL domain (D2), doesn’t change the overall SlpA structure and a functional S-layer is assembled. However, #Cdiff then becomes susceptible to lysozyme, even though the pores are still very narrow (16A). #SlayerStructure #BugSlayers 10/n
Our work revealing the structure and organisation of #BugSlayer in #Cdiff is finally out in @NatComms! Huge team effort from joint 1st authors @PaolaLanzoni, @barwinskasendra, @JSamWilson, @OishikBanerji! #SlayerStructure nature.com/articles/s4146… 1/n
nature.com
Structure and assembly of the S-layer in C. difficile
Nature Communications - The S-layer is a two-dimensional protein array that covers the cell surface of many bacteria and archaea. Here, the authors use high-resolution X-ray crystallography and...
Many new questions to answer: how do things go in/out of the #Cdiff cells? How can we disrupt this armour? How does the #SlayerStructure link to function and infection? Lots to keep us busy, but for now, a huge thank you to the whole @BugS_layers team! 11/11
7. The #Cdiff #BugSlayer is very tightly packed, with pores of only about 11A, so only small ions can diffuse through #SlayerStructure 9/n dlvr.it/SKw0zN #cdi #cdiff
#Cdiff #Slayerstructure #BugSlayer Fascinating story of an arduous journey with happy ending - many congrats Paula and team @paolalanzoni @barwinskasendra
Together with @Joekaryotic, @filipacostavaz, @GillDouce1, @FairweatherNeil, Shauna O’Beirne, Arnaud Basle, Kamel el Omari and Armin Wenger, coordinated by @PerBullough, @RobFagan and @pssalgado! #SlayerStructure 2/n
7. The #Cdiff #BugSlayer is very tightly packed, with pores of only about 11A, so only small ions can diffuse through #SlayerStructure 9/n dlvr.it/SKw0zN #cdi #cdiff
#Cdiff #Slayerstructure #BugSlayer Fascinating story of an arduous journey with happy ending - many congrats Paula and team @paolalanzoni @barwinskasendra
Many new questions to answer: how do things go in/out of the #Cdiff cells? How can we disrupt this armour? How does the #SlayerStructure link to function and infection? Lots to keep us busy, but for now, a huge thank you to the whole @BugS_layers team! 11/11
8. Removing the most exposed SLPL domain (D2), doesn’t change the overall SlpA structure and a functional S-layer is assembled. However, #Cdiff then becomes susceptible to lysozyme, even though the pores are still very narrow (16A). #SlayerStructure #BugSlayers 10/n
7. The #Cdiff #BugSlayer is very tightly packed, with pores of only about 11A, so only small ions can diffuse through #SlayerStructure 9/n

5. The crystal packing of SlpA mimics the assembly of the S-layer in the cells, so interactions in the crystal are a model for how the #BugSlayer assembles. #SlayerStructure 7/n
3. The 3D crystals are made of stacks of 2D lattices of SlpA #xraycrystallography #SlayerStructure #BugSlayer 5/n

2. the complex is maintained by an intricate “paperclip” motif by the interacting domains. #SlayerStructure #xraycrystallography #BugSlayer 4/n

Together with @Joekaryotic, @filipacostavaz, @GillDouce1, @FairweatherNeil, Shauna O’Beirne, Arnaud Basle, Kamel el Omari and Armin Wenger, coordinated by @PerBullough, @RobFagan and @pssalgado! #SlayerStructure 2/n
Our work revealing the structure and organisation of #BugSlayer in #Cdiff is finally out in @NatComms! Huge team effort from joint 1st authors @PaolaLanzoni, @barwinskasendra, @JSamWilson, @OishikBanerji! #SlayerStructure nature.com/articles/s4146… 1/n
nature.com
Structure and assembly of the S-layer in C. difficile
Nature Communications - The S-layer is a two-dimensional protein array that covers the cell surface of many bacteria and archaea. Here, the authors use high-resolution X-ray crystallography and...
3. The 3D crystals are made of stacks of 2D lattices of SlpA #xraycrystallography #SlayerStructure #BugSlayer 5/n

2. the complex is maintained by an intricate “paperclip” motif by the interacting domains. #SlayerStructure #xraycrystallography #BugSlayer 4/n

7. The #Cdiff #BugSlayer is very tightly packed, with pores of only about 11A, so only small ions can diffuse through #SlayerStructure 9/n

Something went wrong.
Something went wrong.
United States Trends
- 1. Auburn 40.2K posts
- 2. Duke 31.9K posts
- 3. Bama 29.3K posts
- 4. Stockton 22.1K posts
- 5. Miami 131K posts
- 6. Ole Miss 38.4K posts
- 7. Lane Kiffin 48.1K posts
- 8. Notre Dame 25.6K posts
- 9. Stanford 9,755 posts
- 10. #SurvivorSeries 186K posts
- 11. Virginia 48.6K posts
- 12. Austin Theory 5,012 posts
- 13. Cam Coleman 1,984 posts
- 14. #Toonami 2,966 posts
- 15. Cooper Flagg 8,120 posts
- 16. Seth 21.6K posts
- 17. ACC Championship 8,330 posts
- 18. The ACC 32K posts
- 19. Ewing 1,220 posts
- 20. #RollTide 6,313 posts