#moleculardynamics search results
How Minor Sequence Changes Enable Mechanistic Diversity in MFS Transporters? An Atomic-Level Rationale for Symport Emergence in NarU #MolecularDynamics pubs.acs.org/doi/10.1021/ac… @diwakarshukla #JCIM Vol66 Issue4 #compchem
Current Status of Molecular Dynamics Simulations of Membrane Permeabilization by Antimicrobial Peptides and Pore-Forming Proteins: A Review #MolecularDynamics pubs.acs.org/doi/10.1021/ac… @Magistrato_Lab #JCIM Vol66 Issue4 #Review
Identification of Potential Multitarget Directed Ligands for Alzheimer’s Disease by Coupling Virtual Screening and Experimental Validation #VirtualScreening #MolecularDynamics pubs.acs.org/doi/10.1021/ac… @Paulina_valhor @CeciliaScorza #JCIM Vol66 Issue5 #PharmaceuticalModeling
DynoPore─A Package to Analyze Molecular Dynamics Trajectories of Confined Liquids #MolecularDynamics pubs.acs.org/doi/10.1021/ac… @chrisewels #JCIM Vol66 Issue3 #compchem
Sriraksha Srinivasan, Daniel Álvarez, Stefano Vanni et al. @LabVanni @unifr develop a fast and easy-to-use protocol based on coarse-grained #moleculardynamics simulations to characterize #lipid binding to lipid transfer proteins. hubs.la/Q02KfsXq0 #lipids #biophysics

Allosteric Insights into TCR–pMHC Dynamics: Understanding the Effects of Melanoma-Associated Epitopes #MolecularDynamics pubs.acs.org/doi/10.1021/ac… #JCIM Vol66 Issue3 #Bioinformatics
Metadynamics Simulation Reveals Allosteric Communication Effects of the Flipping Process of the Atypical DLG Motif in RIPK1 #MolecularDynamics #Allostery pubs.acs.org/doi/10.1021/ac… #JCIM Vol66 Issue3 #compchem
MDIntrinsicDimension: Dimensionality-Based Analysis of Collective Motions in Macromolecules from Molecular Dynamics Trajectories #MolecularDynamics pubs.acs.org/doi/10.1021/ac… #JCIM Vol66 Issue5 #ApplicationNote
Understanding the Kinetic Mechanism of Ligands Stabilizing the RAS–CYPA Interaction #MolecularDynamics pubs.acs.org/doi/10.1021/ac… #JCIM Vol66 Issue3 #PharmaceuticalModeling
Let’s look at the kinetic isotope effect — at the molecular level. 🔎 In Darmstadtium-Hydrogen dimers, the D2H+ cation forms faster and more often than the other cation because the H2+ cation vibrates faster, this study shows: go.aps.org/4gJAlY9 #MolecularDynamics

#research #moleculardynamics #compchem #machinelearning Building Water Models Compatible with Charge Scaling Molecular Dynamics (Jungwirth, @hseara7) - @JPhysChem Letters: doi.org/10.1021/acs.jp…
Improving Protein Structure Determination by Integrating Ensemble-Driven Molecular Dynamics with Chemical Shift-Based Restraints #MolecularDynamics pubs.acs.org/doi/10.1021/ac… #JCIM Vol66 Issue5 #compchem
Unbiased #MolecularDynamics simulations identify lipid binding sites in lipid transfer proteins, say Sriraksha Srinivasan, Daniel Álvarez, Stefano Vanni et al. @LabVanni @unifr hubs.la/Q02Kfhhr0 #lipids #biophysics
Recognition of Coexisting Phases in Model Membranes via an Unsupervised Method #MolecularDynamics pubs.acs.org/doi/10.1021/ac… @YuzhuoDai #JCIM Vol66 Issue3 #compchem
Molecular Dynamics-Enhanced Sampling Reveals Electrofusion Mechanisms and Pathways #MolecularDynamics pubs.acs.org/doi/10.1021/ac… #JCIM Vol66 Issue4 #compchem
Decoding Protein–Membrane Binding Interfaces from Surface-Fingerprint-Based Geometric Deep Learning and Molecular Dynamics Simulations #DeepLearning #MolecularDynamics pubs.acs.org/doi/10.1021/ac… @VanLehnGroup #JCIM Vol66 Issue4 #compchem
Evidence for Epibatidine Binding to the Desensitization Gate in α7 nAChR from Molecular Dynamics Simulations and Cryo-EM #MolecularDynamics #CryoEM pubs.acs.org/doi/10.1021/ac… #JCIM Vol66 Issue3 #Letter
Mapping the Allosteric Landscape of PPARγ: a Markov State Modeling and Energetic Analysis Approach #MolecularDynamics #Allostery pubs.acs.org/doi/10.1021/ac… #JCIM Vol66 Issue2 #compchem
Mechanistic Insights into Nanobody-Based Quenchbody Sensing via Structural Modeling and Molecular Simulations #MolecularDynamics pubs.acs.org/doi/10.1021/ac… @LMSpenkelink @haibo_yu @UOW @MolHorizons @arc_qubic #JCIM Vol66 Issue5 #compchem
Atomistic #moleculardynamics simulations of short #peptides can lead to better understanding more complex molecular systems. Join us to hear more from @daviddesancho 🗓️ 12 May 2026, 15:00 CET ✍️ bioexcel.eu/dx7m #ComputerSimulation #biophysics #proteins

🎙️ Weekly GPCR Simulation Digest (2 papers) 📄 Investigating the electrostatics underlying activation of th… pubmed.ncbi.nlm.nih.gov/41546993/ 📄 Salidroside protects against high-altitude hypoxia-induced k… pubmed.ncbi.nlm.nih.gov/41915656/ #GPCR #MolecularDynamics #DrugDiscovery
🎙️ Weekly GPCR Simulation Digest (2 papers) 📄 Multiple modes of cholesterol translocation in the human Smo… pubmed.ncbi.nlm.nih.gov/41811180/ 📄 A Transferable and Robust Computational Framework for Class … pubmed.ncbi.nlm.nih.gov/41774824/ #GPCR #MolecularDynamics #DrugDiscovery
🎙️ Weekly GPCR Simulation Digest (1 papers) 📄 Specificity and Promiscuity of Phosphoinositide Lipid Intera… pubmed.ncbi.nlm.nih.gov/41782363/ #GPCR #MolecularDynamics #DrugDiscovery
🎙️ Weekly GPCR Simulation Digest (2 papers) 📄 Investigating the electrostatics underlying activation of th… pubmed.ncbi.nlm.nih.gov/41546993/ 📄 Salidroside protects against high-altitude hypoxia-induced k… pubmed.ncbi.nlm.nih.gov/41915656/ #GPCR #MolecularDynamics #DrugDiscovery
🎙️ Weekly GPCR Simulation Digest (2 papers) 📄 Multiple modes of cholesterol translocation in the human Smo… pubmed.ncbi.nlm.nih.gov/41811180/ 📄 A Transferable and Robust Computational Framework for Class … pubmed.ncbi.nlm.nih.gov/41774824/ #GPCR #MolecularDynamics #DrugDiscovery
🎙️ Weekly GPCR Simulation Digest (1 papers) 📄 Specificity and Promiscuity of Phosphoinositide Lipid Intera… pubmed.ncbi.nlm.nih.gov/41782363/ #GPCR #MolecularDynamics #DrugDiscovery
CHARMM-GUI Hybrid ML/MM Builder for Hybrid Machine Learning and Molecular Mechanical Modeling and Simulations #MolecularDynamics #MachineLearning pubs.acs.org/doi/10.1021/ac… #JCIM Vol66 Issue6 #ApplicationNote
plotXVG: Batch Generation of Publication-Quality Graphs from GROMACS Output #MolecularDynamics #GROMACS pubs.acs.org/doi/10.1021/ac… #JCIM Vol66 Issue6 #ApplicationNote
Identification of Potential Multitarget Directed Ligands for Alzheimer’s Disease by Coupling Virtual Screening and Experimental Validation #VirtualScreening #MolecularDynamics pubs.acs.org/doi/10.1021/ac… @Paulina_valhor @CeciliaScorza #JCIM Vol66 Issue5 #PharmaceuticalModeling
Mechanistic Insights into Nanobody-Based Quenchbody Sensing via Structural Modeling and Molecular Simulations #MolecularDynamics pubs.acs.org/doi/10.1021/ac… @LMSpenkelink @haibo_yu @UOW @MolHorizons @arc_qubic #JCIM Vol66 Issue5 #compchem
Join our #webinar tomorrow "Multiscale simulation of biomembranes: shape, structure and cellular function" 🗓️ 14 April 2026, 15:00 CET ✍️ bioexcel.eu/y50u #ComputerSimulation #moleculardynamics
Comparative Binding Analysis by Computational Methods and Quartz Crystal Microbalance: Case of hnRNPA2B1 Protein and Irinotecan Drug #MolecularDynamics #Docking pubs.acs.org/doi/10.1021/ac… #JCIM Vol66 Issue5 #compchem
Protonation State-dependent Interaction of Polycationic Polymyxins with the Pseudomonas aeruginosa Outer Membrane Using Colistin A as a Model #MolecularDynamics pubs.acs.org/doi/10.1021/ac… @mattiamori79 #JCIM Vol66 Issue5 #compchem
Improving Protein Structure Determination by Integrating Ensemble-Driven Molecular Dynamics with Chemical Shift-Based Restraints #MolecularDynamics pubs.acs.org/doi/10.1021/ac… #JCIM Vol66 Issue5 #compchem
EdSr: A Novel End-to-End Approach for State-Space Sampling in Molecular Dynamics Simulation #MolecularDynamics pubs.acs.org/doi/10.1021/ac… #JCIM Vol66 Issue5 #compchem
My GPU is currently chewing through the final 50ns Metadynamics runs. Next step: the physical twin. If you know an academic lab for gene synthesis/assays to collaborate with—or a good CRO to hire—my DMs are open! 📩 #ProteinEngineering #TechBio #MolecularDynamics #OpenMM
on Day 2! Dive into advanced molecular dynamics simulations and explore AI-driven QSAR modeling. Understand molecular behavior over time and leverage AI to predict biological activity—bridging computation with real-world drug discovery challenges. #MolecularDynamics #QSAR
A Neural Time-Series Learning Method for Accelerating Free-Energy Perturbation and Rare-Event Molecular Dynamics Simulations #MolecularDynamics pubs.acs.org/doi/10.1021/ac… #JCIM Vol66 Issue5 #MachineLearning #DeepLearning
🔬 Tool #21: Desmond It is known for its efficiency, accuracy, and integration with modern drug discovery platforms. #Desmond #MolecularDynamics #DrugDiscovery #DecodeScience

Enhancing Coarse-Grained Models through Machine Learning #MachineLearning #moleculardynamics pubs.acs.org/doi/10.1021/ac… @TarakChem @kmerz1 #JCIM Vol64 Issue8 #Editorial

Let’s look at the kinetic isotope effect — at the molecular level. 🔎 In Darmstadtium-Hydrogen dimers, the D2H+ cation forms faster and more often than the other cation because the H2+ cation vibrates faster, this study shows: go.aps.org/4gJAlY9 #MolecularDynamics

Sriraksha Srinivasan, Daniel Álvarez, Stefano Vanni et al. @LabVanni @unifr develop a fast and easy-to-use protocol based on coarse-grained #moleculardynamics simulations to characterize #lipid binding to lipid transfer proteins. hubs.la/Q02KfsXq0 #lipids #biophysics

Using optical spectroscopic techniques and #MolecularDynamics simulations, researchers uncover the mechanism behind the controlled release of #vitamin B1 conjugated with citrate-capped gold #nanoparticles with added calcium ions under physiological pH. 🔗 go.aps.org/3Z93Bjy

#MolecularDynamics Developing and Benchmarking Sulfate and Sulfamate Force Field Parameters via Ab Initio Molecular Dynamics Simulations To Accurately Model Glycosaminoglycan Electrostatic Interactions (@hseara7, @DenysBiriukov) - @JCIM_JCTC: doi.org/10.1021/acs.jc… @IOCBPrague

[Selected Paper] #ProteinProteinInteractions , #MolecularDynamics simulation , #AntitumorAgents Article by Prof. Masaki Kita @NagoyaUniv (Nagoya University) #ProteinProteinInteraction #MDsimulation #OnTheCover #FreeAccess academic.oup.com/bcsj/article/9…
![CSJjournals_jp's tweet image. [Selected Paper]
#ProteinProteinInteractions , #MolecularDynamics simulation , #AntitumorAgents
Article by Prof. Masaki Kita @NagoyaUniv (Nagoya University)
#ProteinProteinInteraction #MDsimulation #OnTheCover #FreeAccess
academic.oup.com/bcsj/article/9…](https://pbs.twimg.com/media/GWr3wRGa8AAMzGK.jpg)
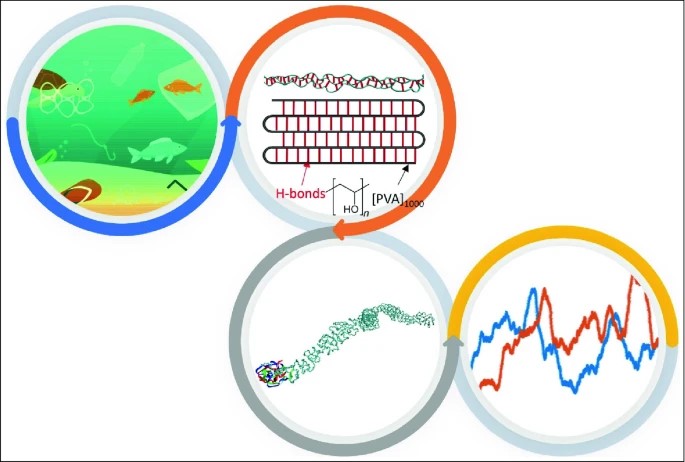
In this #MRSBulletinImpact article, the authors use #moleculardynamics modeling to investigate the large-scale #molecularstructures of a #PVA-#collagen interface and reveal its structure-mechanics relationship. #openaccess loom.ly/18MApr
#research #moleculardynamics #RamanOpticalActivity Detection of Guanine Quadruplexes by Raman Optical Activity and Quantum-Chemical Interpretation of the Spectra (Bouř) - @chemeurj: doi.org/10.1002/chem.2… @IOCBPrague @fzu_avcr @CzechAcademy @matfyz @prfupol

Rucker and Zhang use #MolecularDynamics to investigate inter/intramolecular interactions in lignin, published in #Biomacromolecules! go.acs.org/7A6

#IQFPaper Isothermal compressibility and water self-diffusion coefficient in supercooled salt aqueous solutions using the Madrid-2019 model. Published in Molecular Physics Collaboration with @quimicasUCM #ElectrolyteSolutions #MolecularDynamics bit.ly/4fPdJED

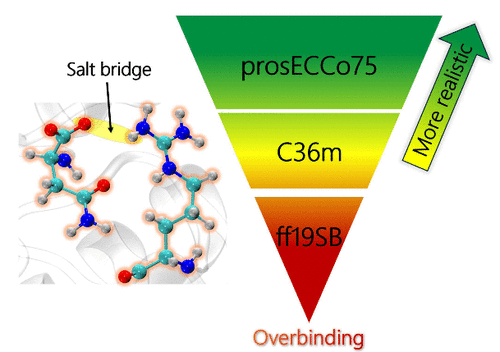
#research #NMR #moleculardynamics #proteins NMR-Derived Salt Bridges in Insulin Analogue: Resolving Artifactual Overbinding in Molecular Dynamics via Charge Scaling (Le Nguyen, Žák, Jungwirth, Lepšík) – J. Phys. Chem. Lett.: doi.org/10.1021/acs.jp… @IOCBPrague @CzechAcademy
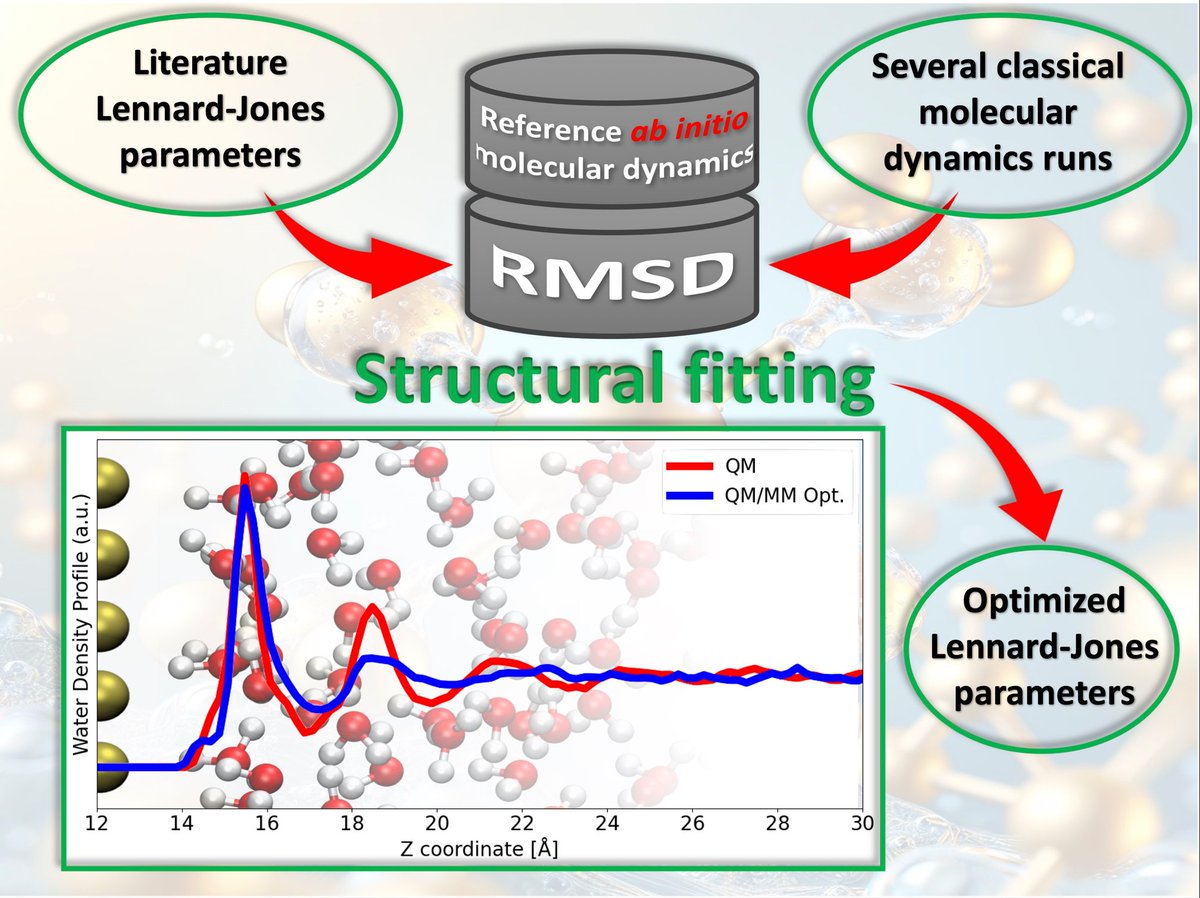
Selected Paper for #MathDay Standard Lennard-Jones parameters often fail at metallic interfaces. Fitting parameters via structural statistical analysis bridges quantum and classical models. Read more: mdpi.com/3696308 #MolecularDynamics #ComputationalChemistry
This joint workshop is suitable for scientists performing or using #biomolecular modelling, #moleculardynamics and/or simulation in their work. Applications close this Sunday 4 June: ebi.ac.uk/training/event… Run in association with @ccpbiosim, @CoSeC_community, & @STFC_Matters

#research #moleculardynamics #NMR #Hyaluronan-#arginine enhanced and dynamic interaction emerges from distinctive molecular signature due to electrostatics and side-chain specificity (Martinez-Seara) - Carbohydr.Polym.: doi.org/10.1016/j.carb… @IOCBPrague @hseara7 @DenysBiriukov

🔬BioExcel-CV19: A breakthrough in understanding #COVID19 proteins & a revolutionary database for #MolecularDynamics simulations. ✍️Daniel Beltrán, Adam Hospital,@jlgelpi Modesto Orozco-@orozco_lab 📰@NAR_Open ➡️shorturl.at/xyNV6 📌DOI: 10.1093/nar/gkad991 #IRBScience
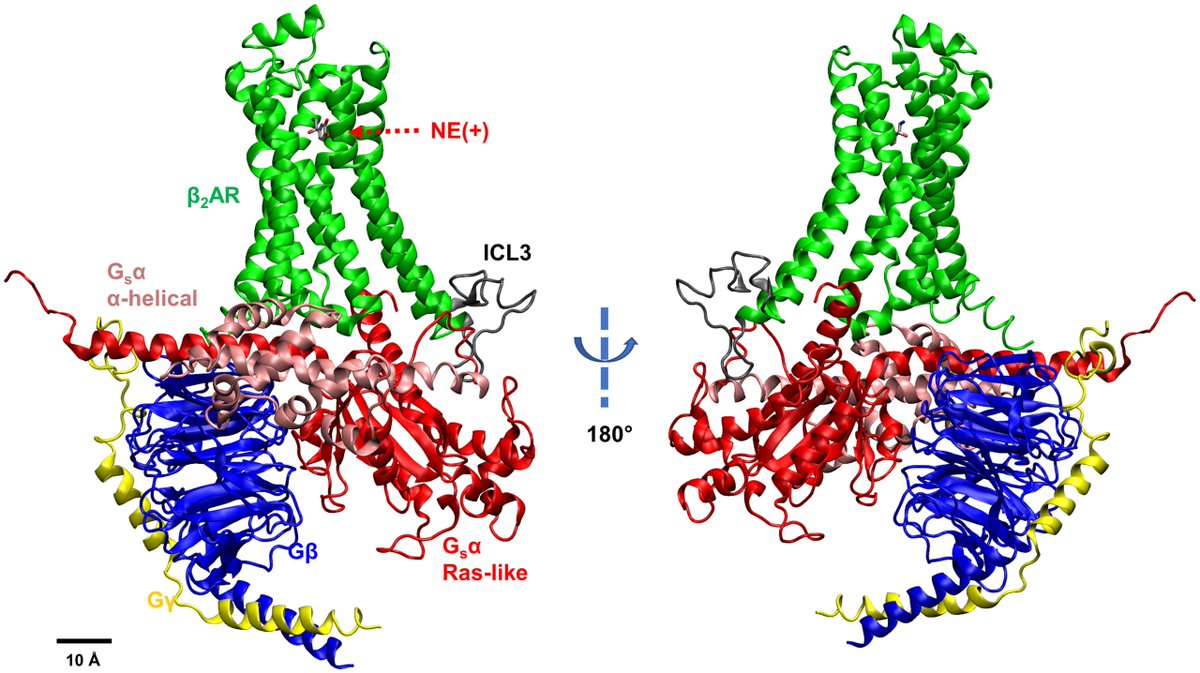
Simulations run on #Anton2 by @ucd_physiology researchers reveal previously unseen activity between G proteins and cell-surface receptor proteins. Results offer clues to designing better G protein-targeting drugs. Read more: bit.ly/3WTVvcv #HPC #MolecularDynamics
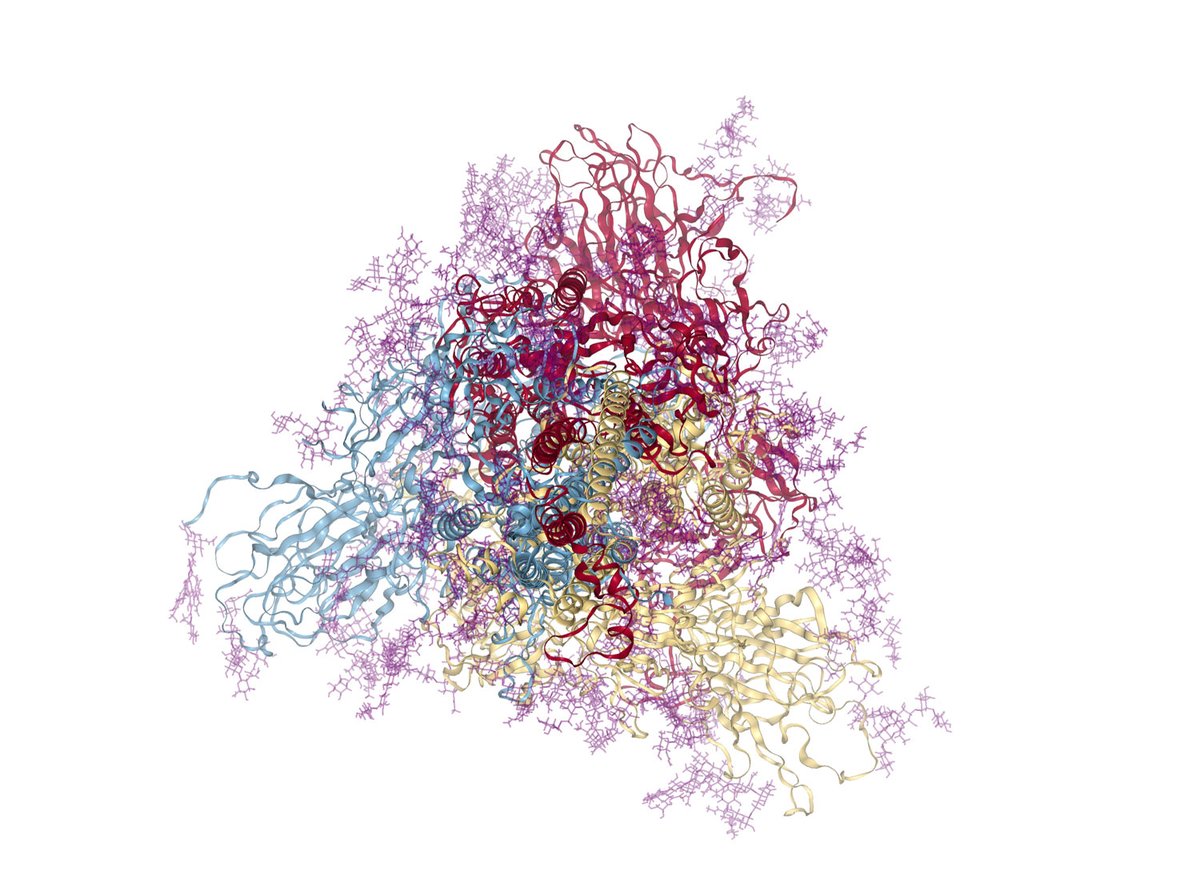
Our very own Thomas Tarenzi presents a fascinating talk on fuzzy interactions on proteins in the yeast acetylation complex at the frontiers in IDP meeting #IDP #moleculardynamics

Espinoza-Arcos, González-Avendaño, Vergara-Jaque, Poblete et al. @UTalca identify a peripheral PIP2-binding site conserved across TRP channels using coarse-grained #MolecularDynamics simulations. hubs.la/Q03F10LN0 #Lipids #ComputationalBiology #IonChannels

We've loved hosting @ccpbiosim this week's workshop on #structuralbioinformatics resources and tools for #moleculardynamics simulations. Run in association with @CoSeC_community and @STFC_Matters. Discover on-demand structural biology training: ebi.ac.uk/training/on-de…

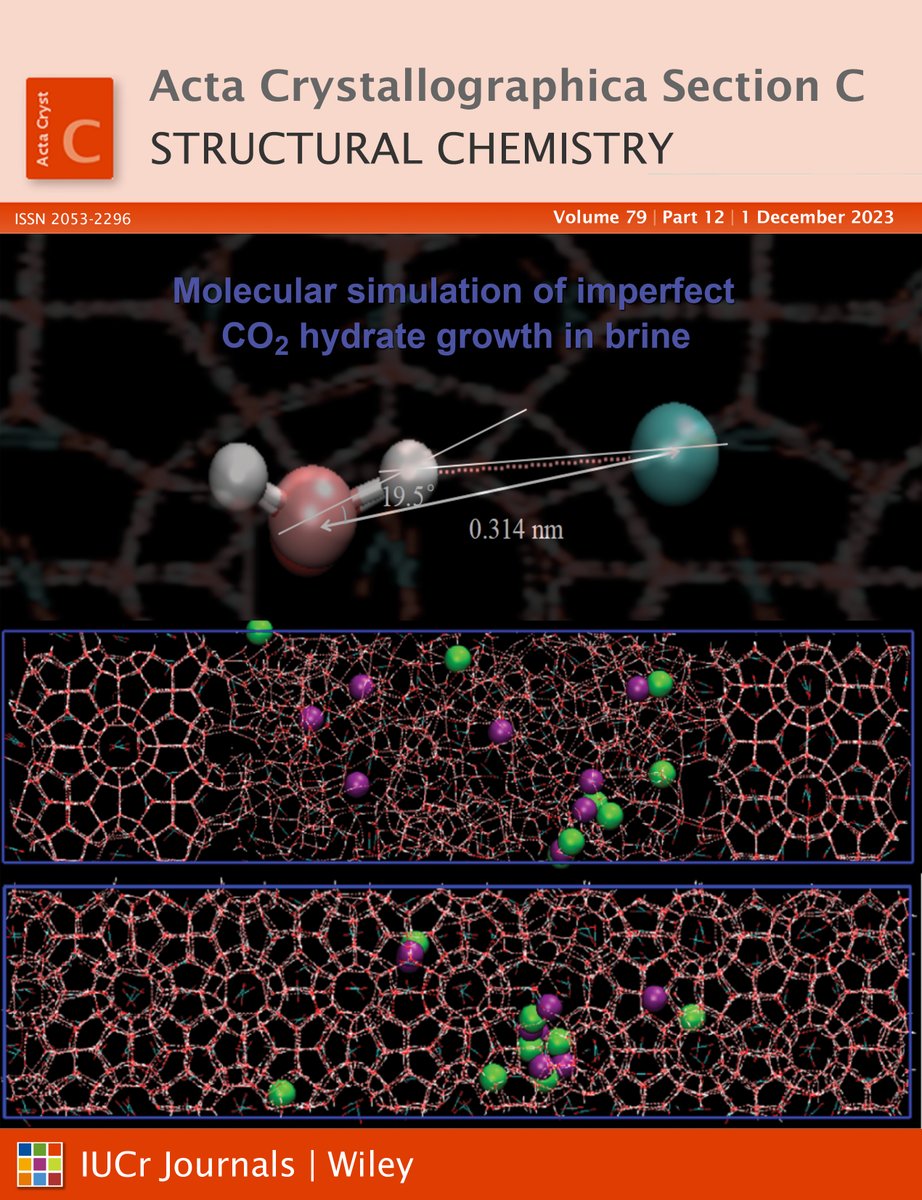
December issue @ActaCrystC The cover article reports the molecular simulation of imperfect structure I CO2 hydrate growth in brine #moleculardynamics #simulation #orderparameter #climatechange #carboncapture #crystallography journals.iucr.org/c/issues/2023/…

Something went wrong.
Something went wrong.
United States Trends
- 1. #AEWDynamite N/A
- 2. #Survivor50 N/A
- 3. Boldy N/A
- 4. Benn N/A
- 5. #mnwild N/A
- 6. Flyers N/A
- 7. Leftover N/A
- 8. #CAGovDebate N/A
- 9. #IgniteTheOrange N/A
- 10. Rust N/A
- 11. Crosby N/A
- 12. Penguins N/A
- 13. Pens N/A
- 14. Luke Weaver N/A
- 15. Bo Jackson N/A
- 16. Duchene N/A
- 17. Doris Burke N/A
- 18. Vientos N/A
- 19. Ciampa N/A
- 20. Grapefruit N/A