#spatialtranscriptomics search results
🚨New #SpatialTranscriptomics #Bioinformatics data resource out in @naturemethods. SODB, a platform with >2,400 manually curated spatial experiments from >25 spatial omics technologies & interactive analytical modules. This🧵will walk you through all the features of SODB [1/33]
![simocristea's tweet image. 🚨New #SpatialTranscriptomics #Bioinformatics data resource out in @naturemethods.
SODB, a platform with >2,400 manually curated spatial experiments from >25 spatial omics technologies & interactive analytical modules.
This🧵will walk you through all the features of SODB [1/33]](https://pbs.twimg.com/media/FpqfK6HaUAAQZAo.jpg)
Ruochen Dong, @linheng_li lab, gave an outstanding presentation on using #SpatialTranscriptomics to interrogate HSC-niche interaction in the mouse fetal liver at #ISSCR2023. He also received merit and travel awards. Congratulations! @ISSCR

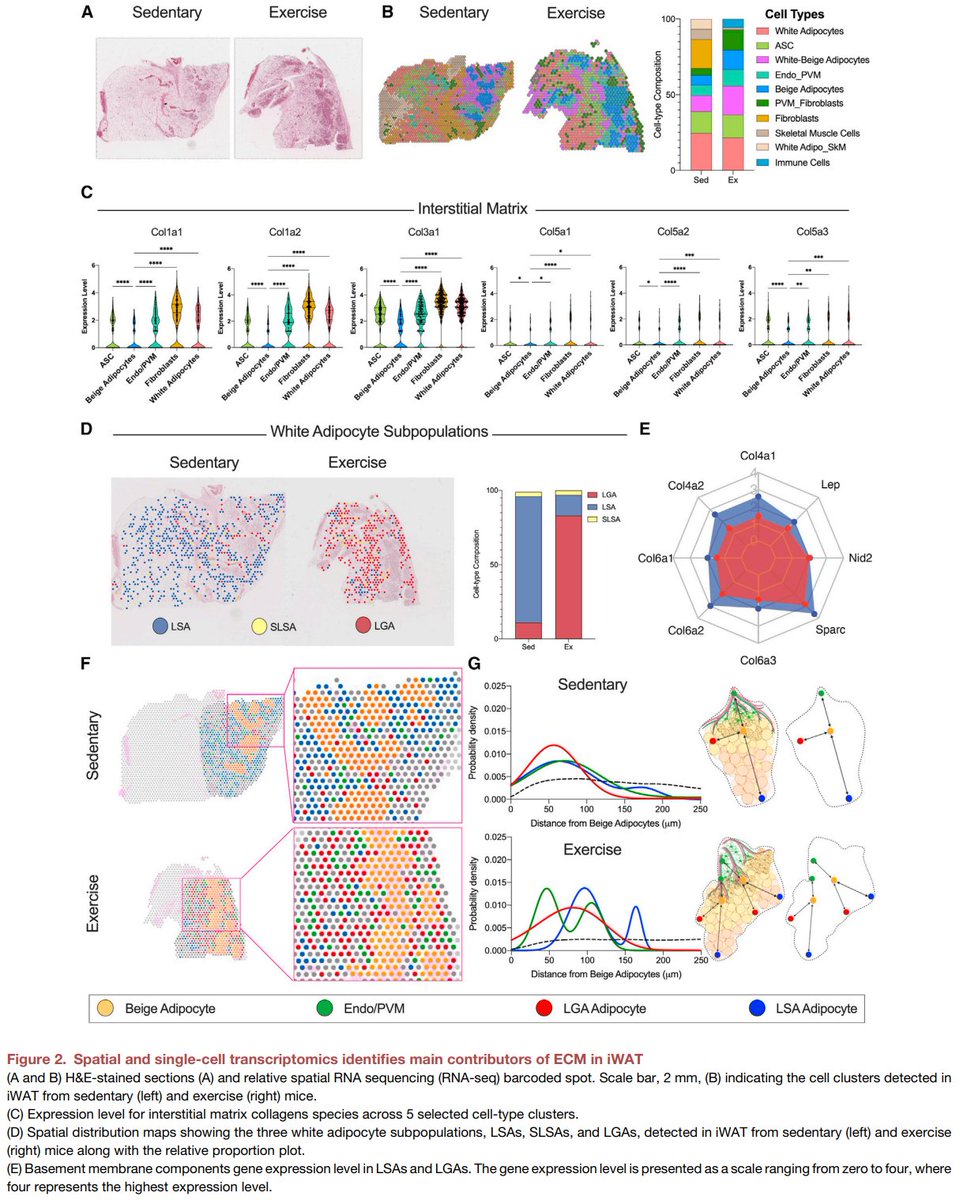
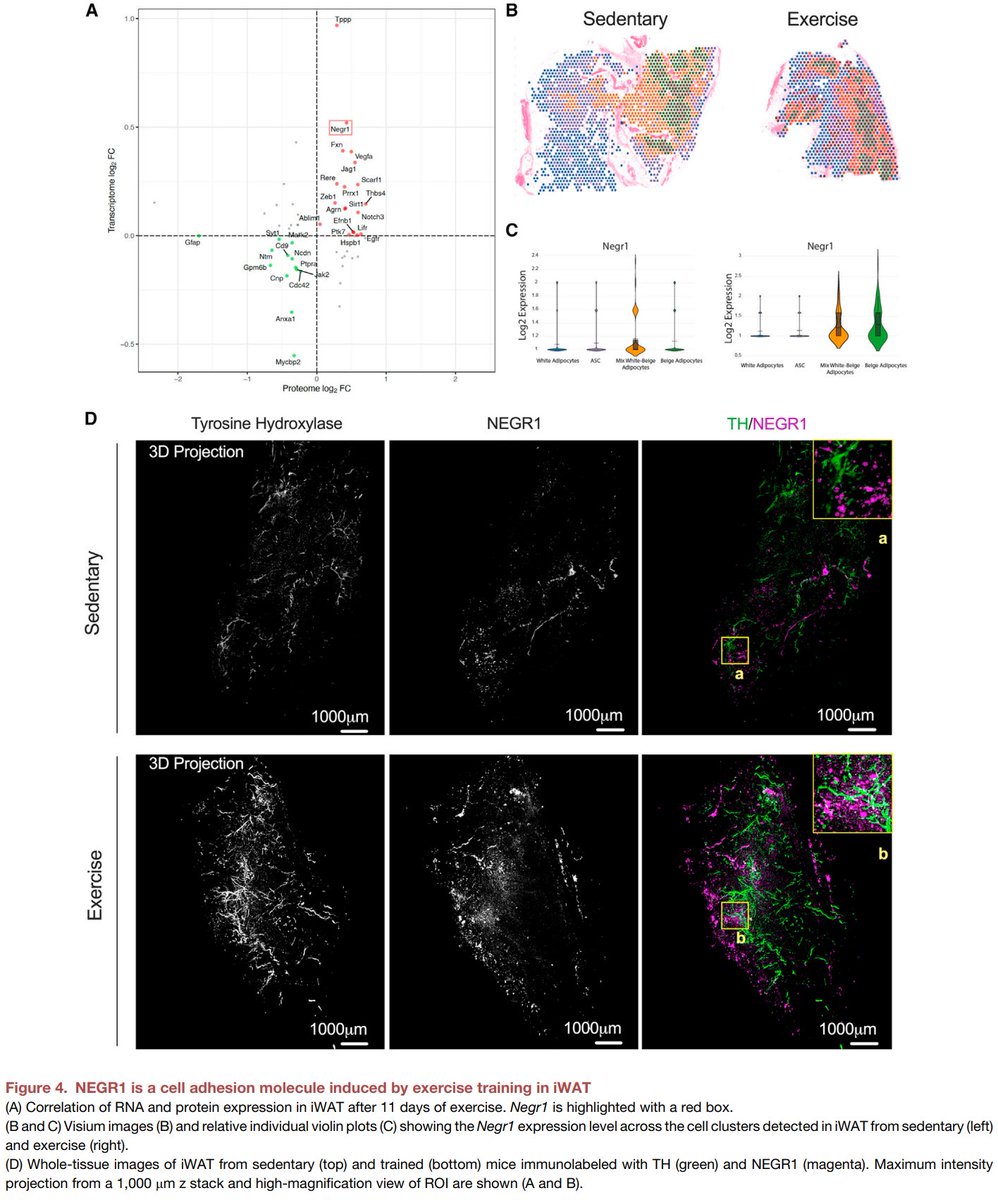
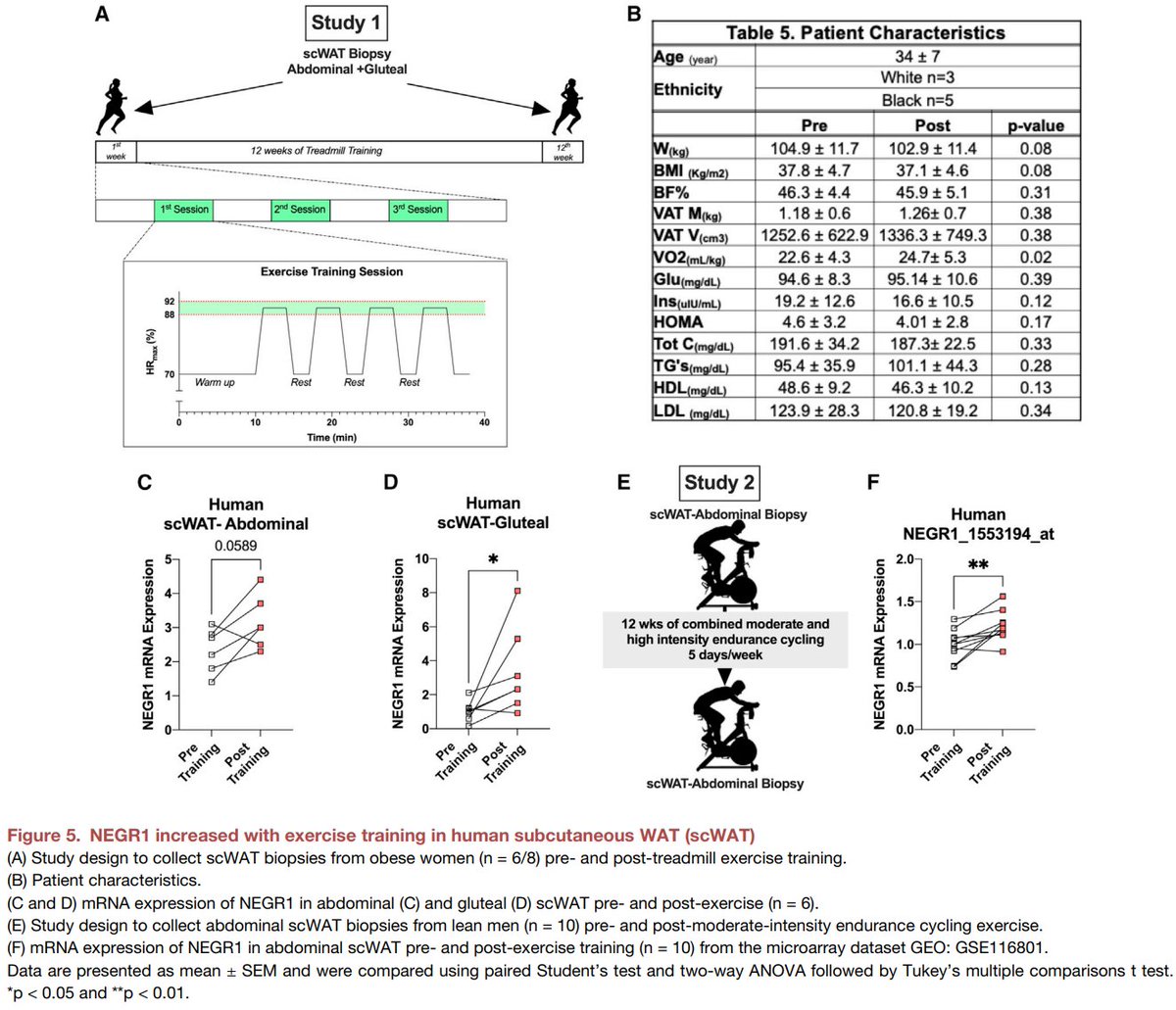
Excited to share our #SingleCell and #SpatialTranscriptomics analysis of #ExerciseTraining remodeling of #WhiteAdiposeTissue through innervation, vascularization, and #ExtracellularMatrix in mouse and human with #LaurieGoodyear #JanWillemMiddelbeek @PasqualeNigro9 @MVamvini…
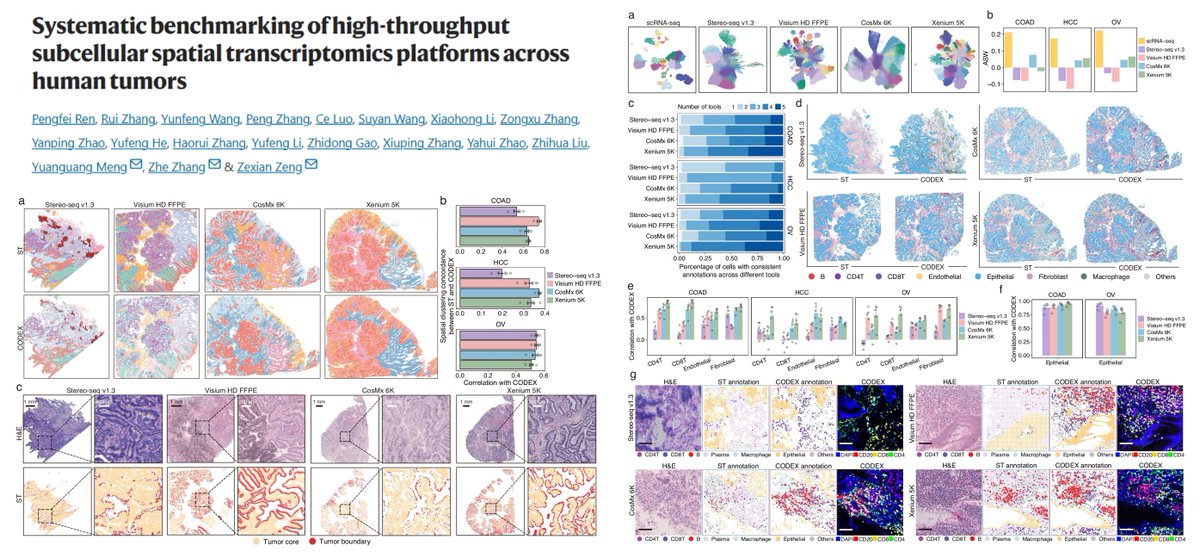
Systematic comparisons among 4 subcellular-resolution #SpatialTranscriptomics methods Stereo-seq v1.3 Visium HD FFPE CosMx 6k Xenium 5k Human Tumors Ground truth CODEX, +scRNAseq Gene detection sensitivity (CosMx performance...🙁) (Spatial) false positives Transcript-protein…
How do we uncover cell-cell interactions from #SpatialTranscriptomics data? Excited to share Niche-DE (doi.org/10.1186/s13059…). Niche-DE asks how does a cell’s transcriptome depend on its spatial neighbors, by identifying Niche-Differentially Expressed genes. 1/9

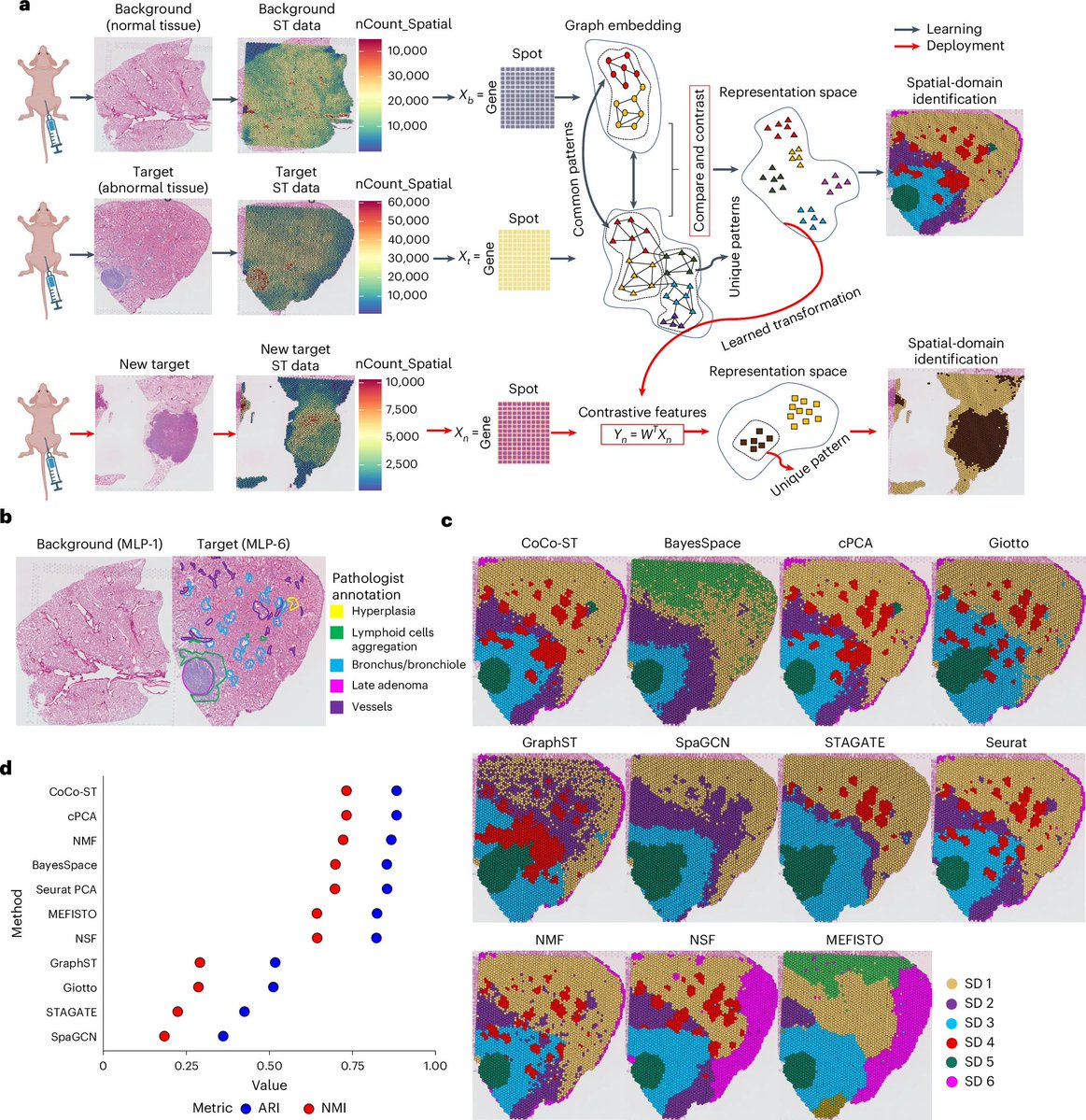
CoCo-ST detects global and local biological structures in spatial transcriptomics datasets. #SpatialTranscriptomics #VarianceIdentification @NatureCellBio nature.com/articles/s4155…
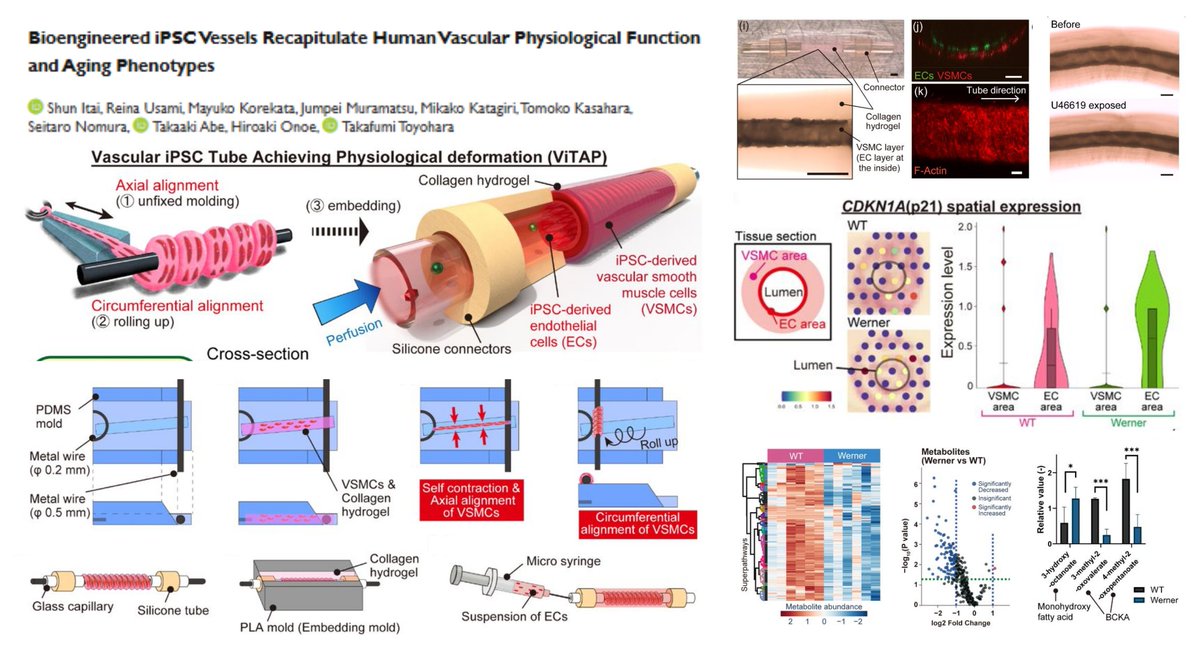
#WernerSyndrome hiPSC-derived engineered vessel #SpatialTranscriptomics #Visium @10xGenomics + #Metabolomics▶️ #BranchedChainKetoAcid⬇️ #MonohydroxyFattyAcid⬆️ in Werner vessels with traits of #PrematureAging Aligned #SmoothMuscleCell in Collagen hydrogel➡️ Roll-up around φ…
Comparing single-cell #SpatialTranscriptomics methods on FFPE human tumor #TissueMicroArray CosMx 1k vs MERFISH 500 vs Xenium-UM/-MM 339 93 common genes lung adenocarcinoma pleural mesothelioma Corr. Bulk RNAseq, GeoMx "better F1-scores with Xenium-MM (median, > 75%) than…

We are very very VERY excited to be running #cosmx #singlecell #spatialtranscriptomics experiments in @UofGlasgow @UofGCancerSci @UofGMVLS Well done to the @LabSpatialNBJ team @nanostringtech


Excellent talk from @Sandy_Figiel in @Uroweb #EAUlab session at #EAU25 presenting our cool data (I'm bias!) on #SpatialTranscriptomics & #ClonalAnalysis to detect unique features of metastasing clones in #ProstateCancer. @OxPCaBiol #SPACE_Study #3DLightsheet




OmiCLIP #Visualomics github.com/GuangyuWangLab… An amazing Histology (HE) - #SpatialTranscriptomics foundation model Contrastive Learning: Image encoder + Gene/Text encoder Not Generative yet Building an ST-bank of 2.2 M image-Visium pairs in 1007 tissue samples, 32 organs Work…

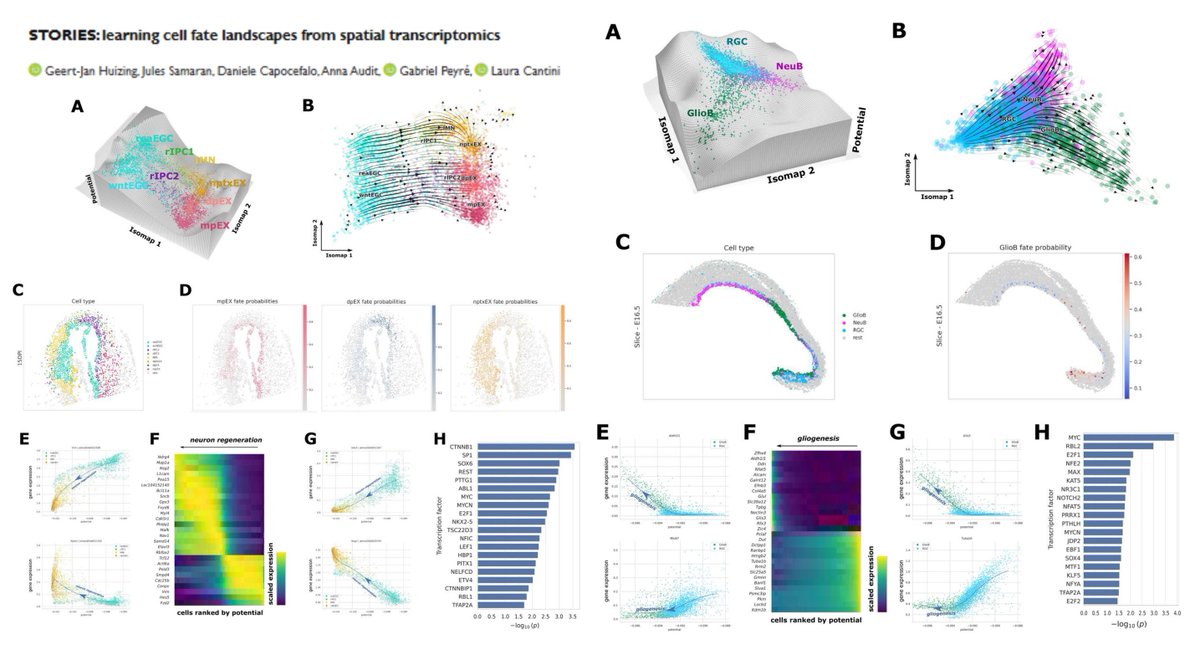
STORIES SpatioTemporal Omics eneRgIES Learning Waddington epigenetic (+regulon) landscape from multi-time-point Stereo-seq #SpatialTranscriptomics Potential-based Fused Gromov-Wasserstein #OptimalTransport Linear▶️gene expression error Quadratic▶️spatial consistency @gjhuizing…
🔬 A new computational approach pioneered by researchers at the @broadinstitute eliminates time-intensive imaging, enabling high-resolution spatial mapping of gene expression in tissue. See how they're making #SpatialTranscriptomics easier for everyone: bit.ly/3G9Hrau
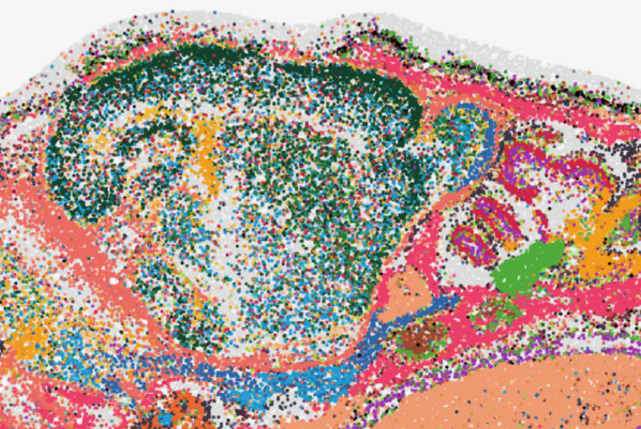

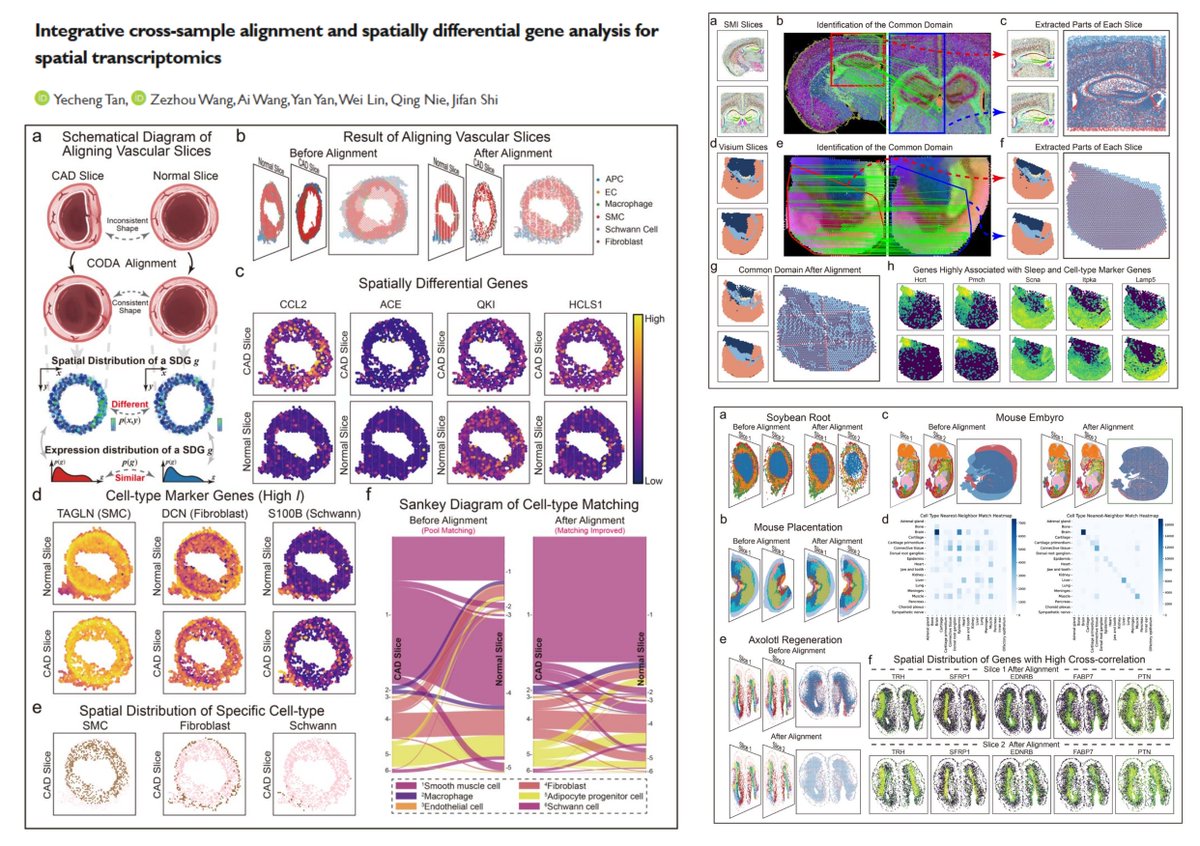
CODA integrative CrOss-sample alignment and spatially Differential gene Analysis for #SpatialTranscriptomics Global rigid alignment➡️ Shared coordinate grid embedding➡️ 3-channel image representation➡️ LightGlue common domain identification➡️ Local nonlinear alignment Work with…
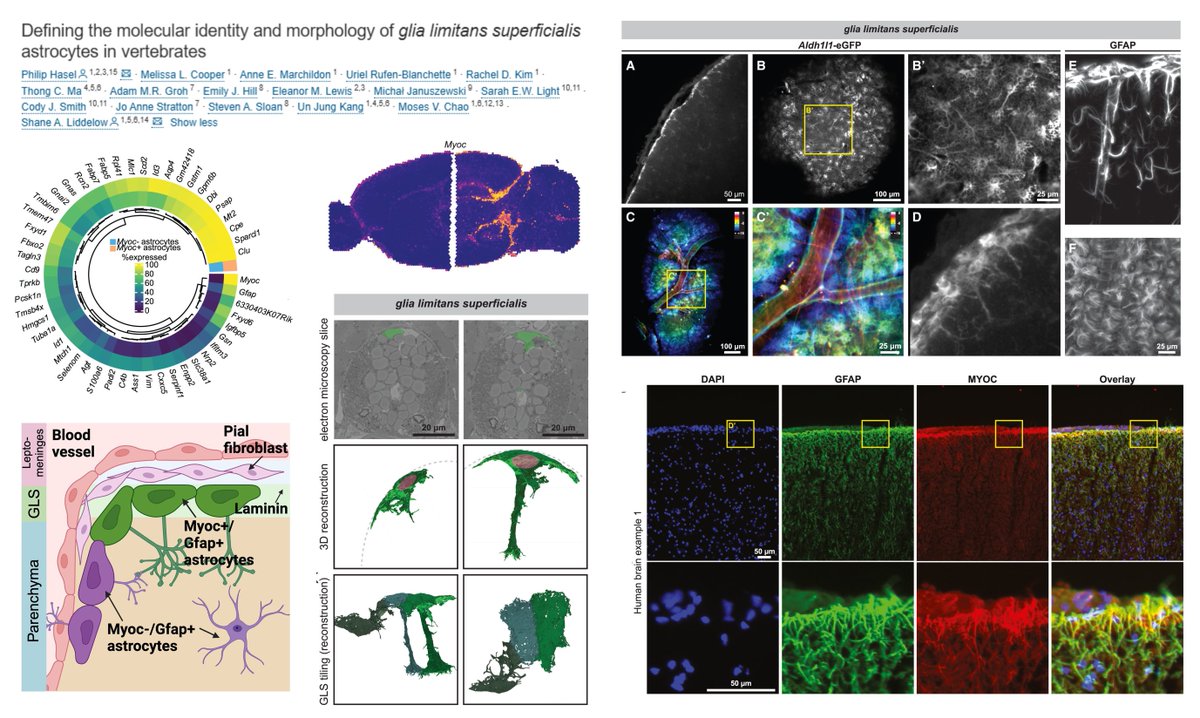
Myocilin marks the glia limitans superficialis Astrocyte Re-analysis of brain-wide #SpatialTranscriptomics & sc-/sn-RNAseq data➡️ Myoc+ 2% of total astrocytes that cover brain/spinal cord surface and extend processes into parenchyma Any marker for glia limitans perivascularis…
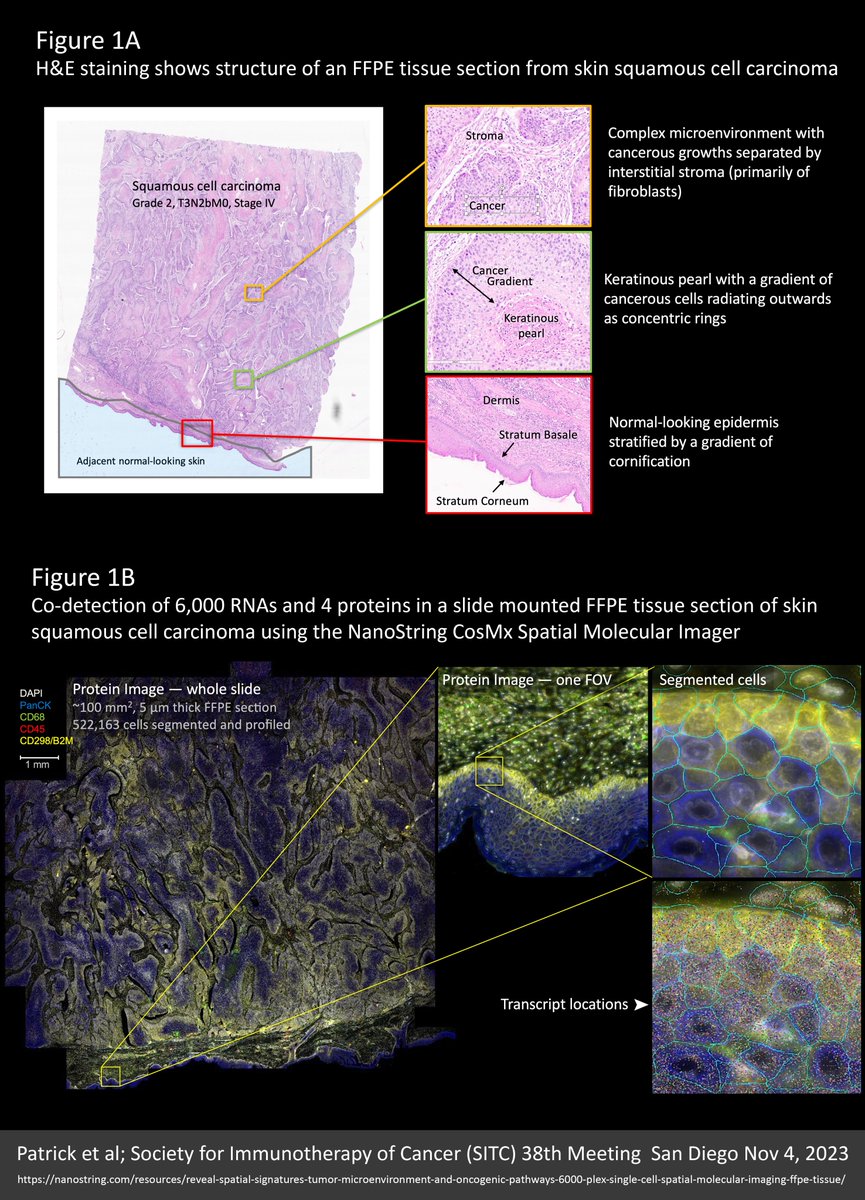
We’re all wondering what the future of #spatialtranscriptomics (ST) has in store. Well, I think the near future will look a lot like the new 6000-plex dataset reviewed here (link to the original source at the end of this post). SAMPLE An entire 100 mm² intact 5 µm FFPE section…



Pleased to share our latest #SpatialTranscriptomics output now on @biorxivpreprint We show marked variation in gene expr relating to @Decipher_VCYT, #Oncotype_DX & #ProstaDiag genomic scores Being presented by @Sandy_Figiel this afternoon #EAU23!! biorxiv.org/content/10.110…
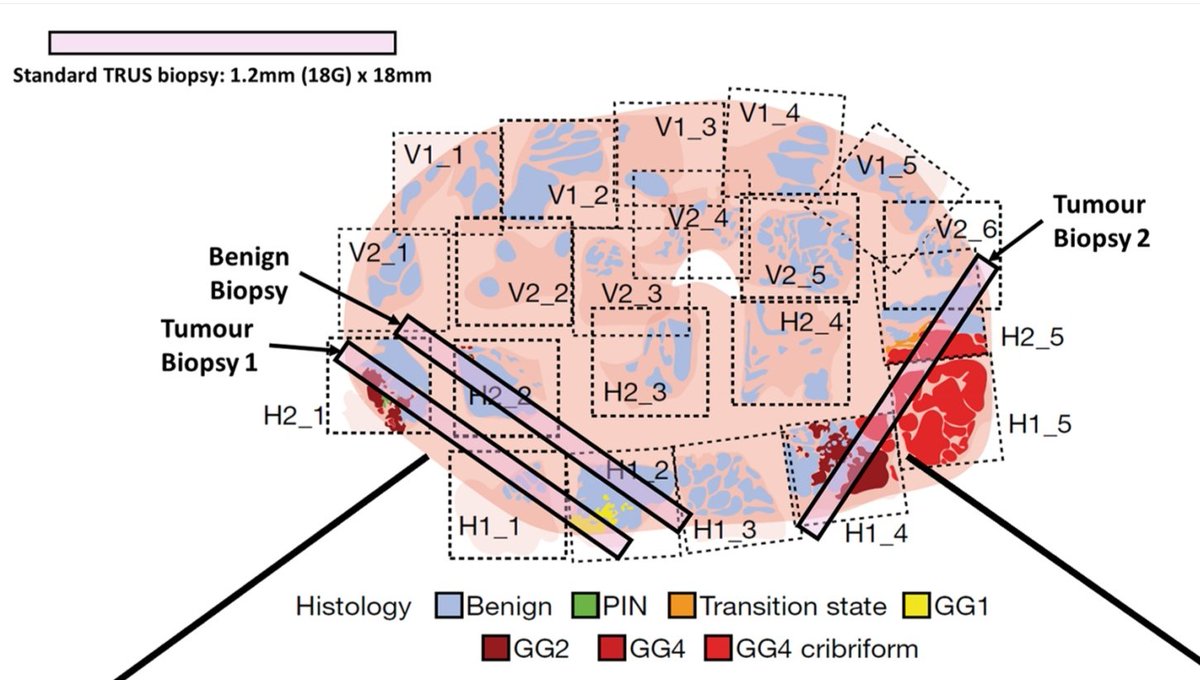



"We all love the pretty pictures we get from #SpatialTranscriptomics but the real value lies in the ability to precisely determine the cellular composition & cell-cell relationships - not possible with bulk RNA or even scRNA approaches" 💯% well said @bayraktar_lab! 👏 #FoG2025




Something went wrong.
Something went wrong.
United States Trends
- 1. #SmackDown 31.8K posts
- 2. Tulane 9,764 posts
- 3. Gunther 20K posts
- 4. North Texas 6,765 posts
- 5. LA Knight 10.3K posts
- 6. #ROHFinalBattle 15K posts
- 7. #OPLive 2,489 posts
- 8. Anthony Davis 1,560 posts
- 9. #TNAFinalResolution 5,614 posts
- 10. Cocona 26.1K posts
- 11. Kennesaw State 3,376 posts
- 12. Mark Pope 3,114 posts
- 13. UNLV 3,629 posts
- 14. Boise State 2,430 posts
- 15. Trouba N/A
- 16. #OPNation N/A
- 17. Jimmy Rogers 2,407 posts
- 18. Wes Miller N/A
- 19. Jalen Johnson 5,064 posts
- 20. John Cena 31.7K posts